7YHO
 
 | |
7YHP
 
 | |
2R3N
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | 3-cyclopropyl-5-phenyl-N-(pyridin-3-ylmethyl)pyrazolo[1,5-a]pyrimidin-7-amine, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.63 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2R3G
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | 6-(2-fluorophenyl)-N-(pyridin-3-ylmethyl)imidazo[1,2-a]pyrazin-8-amine, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.55 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2HXA
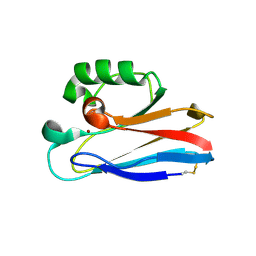 
 | |
2HX9
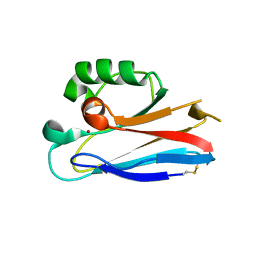 
 | |
2HX7
 
 | |
2HX8
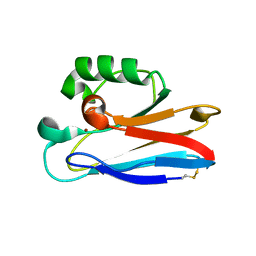 
 | |
2R3J
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | 3-bromo-5-phenyl-N-(pyridin-3-ylmethyl)pyrazolo[1,5-a]pyrimidin-7-amine, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2R3H
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | 3-methyl-N-(pyridin-4-ylmethyl)imidazo[1,2-a]pyrazin-8-amine, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2R3L
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | 3-bromo-6-phenyl-N-(pyrimidin-5-ylmethyl)imidazo[1,2-a]pyridin-8-amine, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.65 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2R3O
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | (5-phenyl-7-(pyridin-3-ylmethylamino)pyrazolo[1,5-a]pyrimidin-3-yl)methanol, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.8 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2R3I
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | 5-(2-fluorophenyl)-N-(pyridin-4-ylmethyl)pyrazolo[1,5-a]pyrimidin-7-amine, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.28 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2R3M
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | Cell division protein kinase 2, N-((2-aminopyrimidin-5-yl)methyl)-5-(2,6-difluorophenyl)-3-ethylpyrazolo[1,5-a]pyrimidin-7-amine | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
2R3F
 
 | | Crystal Structure of Cyclin-Dependent Kinase 2 with inhibitor | | 分子名称: | 5-(2,3-dichlorophenyl)-N-(pyridin-4-ylmethyl)pyrazolo[1,5-a]pyrimidin-7-amine, Cell division protein kinase 2 | | 著者 | Fischmann, T.O, Hruza, A.W, Madison, V.M, Duca, J.S. | | 登録日 | 2007-08-29 | | 公開日 | 2008-01-22 | | 最終更新日 | 2011-07-13 | | 実験手法 | X-RAY DIFFRACTION (1.5 Å) | | 主引用文献 | Structure-guided discovery of cyclin-dependent kinase inhibitors.
Biopolymers, 89, 2008
|
|
7WOF
 
 | | SARS-CoV-2 3CLpro | | 分子名称: | (2S,3S)-3-methyl-N-[(2S)-1-oxidanylidene-3-[(3S)-2-oxidanylidenepiperidin-3-yl]propan-2-yl]-2-[[(E)-3-phenylprop-2-enoyl]amino]pentanamide, 3C-like proteinase | | 著者 | Wang, Y, Ye, S. | | 登録日 | 2022-01-21 | | 公開日 | 2022-04-13 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.72 Å) | | 主引用文献 | Discovery of SARS-CoV-2 3CL Pro Peptidomimetic Inhibitors through the Catalytic Dyad Histidine-Specific Protein-Ligand Interactions.
Int J Mol Sci, 23, 2022
|
|
7WOH
 
 | | SARS-CoV-2 3CLpro | | 分子名称: | (2S)-4-methyl-N-[(2S)-1-oxidanylidene-3-[(3S)-2-oxidanylidenepiperidin-3-yl]propan-2-yl]-2-[[(2S)-3-phenyl-2-[[(E)-3-phenylprop-2-enoyl]amino]propanoyl]amino]pentanamide, 3C-like proteinase | | 著者 | Wang, Y, Ye, S. | | 登録日 | 2022-01-21 | | 公開日 | 2022-04-13 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.72 Å) | | 主引用文献 | Discovery of SARS-CoV-2 3CL Pro Peptidomimetic Inhibitors through the Catalytic Dyad Histidine-Specific Protein-Ligand Interactions.
Int J Mol Sci, 23, 2022
|
|
7WO1
 
 | |
7WO3
 
 | | SARS-CoV-2 3CLpro | | 分子名称: | (2S)-2-[[(2S)-2-[[(E)-3-(4-methoxyphenyl)prop-2-enoyl]amino]-3-methyl-butanoyl]amino]-4-methyl-N-[(2S)-1-oxidanylidene-3-[(3S)-2-oxidanylidenepiperidin-3-yl]propan-2-yl]pentanamide, 3C-like proteinase | | 著者 | Wang, Y, Ye, S. | | 登録日 | 2022-01-20 | | 公開日 | 2022-04-13 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (2.01 Å) | | 主引用文献 | Discovery of SARS-CoV-2 3CL Pro Peptidomimetic Inhibitors through the Catalytic Dyad Histidine-Specific Protein-Ligand Interactions.
Int J Mol Sci, 23, 2022
|
|
7WO2
 
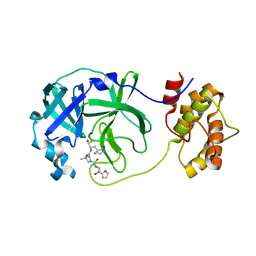 | | SARS-CoV-2 3CLPro Peptidomimetic Inhibitor TPM5 | | 分子名称: | 3C-like proteinase, N-[(2S)-3-methyl-1-[[(2S)-4-methyl-1-oxidanylidene-1-[[(2S)-1-oxidanylidene-3-[(3S}-2-oxidanylidenepiperidin-3-yl]propan-2-yl]amino]pentan-2-yl]amino]-1-oxidanylidene-butan-2-yl]furan-2-carboxamide | | 著者 | Wang, Y, Ye, S. | | 登録日 | 2022-01-20 | | 公開日 | 2022-12-28 | | 最終更新日 | 2023-11-29 | | 実験手法 | X-RAY DIFFRACTION (1.96 Å) | | 主引用文献 | Discovery of SARS-CoV-2 3CL Pro Peptidomimetic Inhibitors through the Catalytic Dyad Histidine-Specific Protein-Ligand Interactions.
Int J Mol Sci, 23, 2022
|
|
7Y8O
 
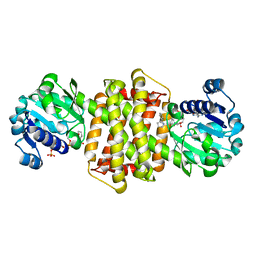 | | Structure of ScIRED-R3-V4 from Streptomyces clavuligerus in complex with 5-(3-fluorophenyl)-3,4-dihydro-2H-pyrrole | | 分子名称: | 2-[2,5-bis(fluoranyl)phenyl]pyrrolidine, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, SciR | | 著者 | Zhang, L.L, Liu, W.D, Shi, M, Huang, J.W, Yang, Y, Chen, C.C, Guo, R.T. | | 登録日 | 2022-06-24 | | 公開日 | 2023-03-22 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Structure of ScIRED-R3-V4 from Streptomyces clavuligerus in complex with 5-(3-fluorophenyl)-3,4-dihydro-2H-pyrrole
to be published
|
|
3E85
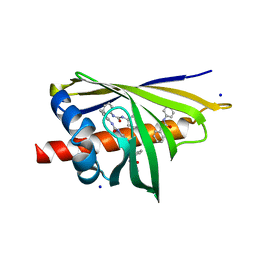 
 | | Crystal Structure of Pathogenesis-related Protein LlPR-10.2B from yellow lupine in complex with Diphenylurea | | 分子名称: | 1,3-DIPHENYLUREA, PR10.2B, SODIUM ION | | 著者 | Fernandes, H.C, Bujacz, G, Bujacz, A, Sikorski, M.M, Jaskolski, M. | | 登録日 | 2008-08-19 | | 公開日 | 2009-03-03 | | 最終更新日 | 2023-08-30 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Cytokinin-induced structural adaptability of a Lupinus luteus PR-10 protein.
Febs J., 276, 2009
|
|
1LHZ
 
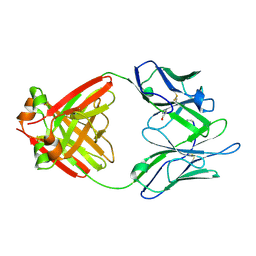 | | Structure of a Human Bence-Jones Dimer Crystallized in U.S. Space Shuttle Mission STS-95: 293K | | 分子名称: | IMMUNOGLOBULIN LAMBDA LIGHT CHAIN | | 著者 | Terzyan, S.S, DeWitt, C.R, Ramsland, P.A, Bourne, P.C, Edmundson, A.B. | | 登録日 | 2002-04-17 | | 公開日 | 2003-07-01 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (2.3 Å) | | 主引用文献 | Comparison of the three-dimensional structures of a human Bence-Jones dimer
crystallized on Earth and aboard US Space Shuttle Mission STS-95
J.MOL.RECOG., 16, 2003
|
|
1LGV
 
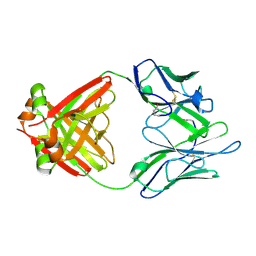 | | Structure of a Human Bence-Jones Dimer Crystallized in U.S. Space Shuttle Mission STS-95: 100K | | 分子名称: | IMMUNOGLOBULIN LAMBDA LIGHT CHAIN | | 著者 | Terzyan, S.S, DeWitt, C.R, Ramsland, P.A, Bourne, P.C, Edmundson, A.B. | | 登録日 | 2002-04-16 | | 公開日 | 2003-07-01 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (1.95 Å) | | 主引用文献 | Comparison of the three-dimensional structures of a human Bence-Jones dimer
crystallized on Earth and aboard US Space Shuttle Mission STS-95
J.MOL.RECOG., 16, 2003
|
|
1KTW
 
 | | IOTA-CARRAGEENASE COMPLEXED TO IOTA-CARRAGEENAN FRAGMENTS | | 分子名称: | 3,6-anhydro-2-O-sulfo-alpha-D-galactopyranose-(1-3)-4-O-sulfo-beta-D-galactopyranose, 3,6-anhydro-2-O-sulfo-alpha-D-galactopyranose-(1-3)-4-O-sulfo-beta-D-galactopyranose-(1-4)-3,6-anhydro-2-O-sulfo-alpha-D-galactopyranose-(1-3)-4-O-sulfo-beta-D-galactopyranose, CALCIUM ION, ... | | 著者 | Michel, G, Kahn, R, Dideberg, O. | | 登録日 | 2002-01-18 | | 公開日 | 2003-06-10 | | 最終更新日 | 2023-08-16 | | 実験手法 | X-RAY DIFFRACTION (2 Å) | | 主引用文献 | The Structural Bases of the Processive Degradation of iota-Carrageenan, a Main Cell Wall Polysaccharide of Red Algae.
J.Mol.Biol., 334, 2003
|
|
