8EUJ
 
 | | Class2 of the INO80-Nucleosome complex | | 分子名称: | DNA (147-MER), Histone H2A type 1, Histone H2B 1.1, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Narlikar, G, Cheng, Y.F. | | 登録日 | 2022-10-18 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-05-29 | | 実験手法 | ELECTRON MICROSCOPY (3.36 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8ETT
 
 | | Class1 of the INO80-Hexasome complex | | 分子名称: | DNA (110-MER), Histone H2A type 1, Histone H2B 1.1, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Cheng, Y.F, Narlikar, G. | | 登録日 | 2022-10-17 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-04-03 | | 実験手法 | ELECTRON MICROSCOPY (6.68 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8ETV
 
 | | Class2 of the INO80-Hexasome complex | | 分子名称: | DNA (110-MER), Histone H2A type 1, Histone H2B 1.1, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Cheng, Y.F, Narlikar, G. | | 登録日 | 2022-10-17 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-04-03 | | 実験手法 | ELECTRON MICROSCOPY (3.16 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8EU2
 
 | | Class3 of the INO80-Hexasome complex | | 分子名称: | DNA (110-MER), Histone H2A type 1, Histone H2B 1.1, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Cheng, Y.F, Narlikar, G. | | 登録日 | 2022-10-18 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-04-03 | | 実験手法 | ELECTRON MICROSCOPY (2.93 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8EUE
 
 | | Class1 of the INO80-Nucleosome complex | | 分子名称: | DNA (147-MER), Histone H2A type 1, Histone H2B 1.1, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Narlikar, G, Cheng, Y.F. | | 登録日 | 2022-10-18 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-04-03 | | 実験手法 | ELECTRON MICROSCOPY (3.48 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8EU9
 
 | | Class1 of the INO80-Nucleosome complex | | 分子名称: | ADENOSINE-5'-DIPHOSPHATE, Actin-related protein 5, Chromatin-remodeling ATPase INO80, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Cheng, Y.F, Narlikar, G. | | 登録日 | 2022-10-18 | | 公開日 | 2023-07-12 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.48 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8ETU
 
 | | Class2 of the INO80-Hexasome complex | | 分子名称: | ADENOSINE-5'-DIPHOSPHATE, Actin-related protein 5, Chromatin-remodeling ATPase INO80, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Cheng, Y.F, Narlikar, G. | | 登録日 | 2022-10-17 | | 公開日 | 2023-07-19 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (2.8 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8ETW
 
 | | Class3 of INO80-Hexasome complex | | 分子名称: | ADENOSINE-5'-DIPHOSPHATE, Actin-related protein 5, Chromatin-remodeling ATPase INO80, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Narlikar, G, Cheng, Y.F. | | 登録日 | 2022-10-17 | | 公開日 | 2023-07-19 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (2.64 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
8ETS
 
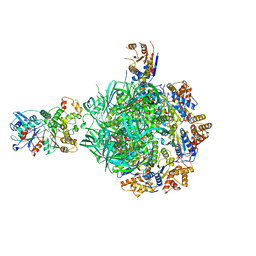 | | Class1 of the INO80-Hexasome complex | | 分子名称: | ADENOSINE-5'-DIPHOSPHATE, Actin-related protein 5, Chromatin-remodeling ATPase INO80, ... | | 著者 | Wu, H, Munoz, E, Gourdet, M, Cheng, Y.F, Narlikar, G. | | 登録日 | 2022-10-17 | | 公開日 | 2023-07-19 | | 最終更新日 | 2024-06-19 | | 実験手法 | ELECTRON MICROSCOPY (3.04 Å) | | 主引用文献 | Reorientation of INO80 on hexasomes reveals basis for mechanistic versatility.
Science, 381, 2023
|
|
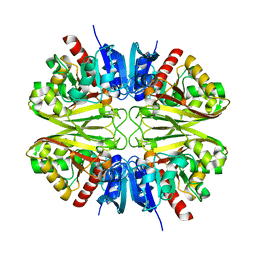
8G16
 
 | |
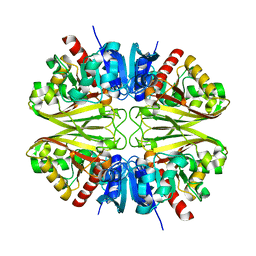
8G15
 
 | |
8G17
 
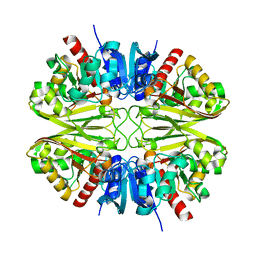 | | CryoEM structure of wild-type GAPDH | | 分子名称: | Glyceraldehyde-3-phosphate dehydrogenase | | 著者 | Choi, W.Y, Wu, H, Cheng, Y.F. | | 登録日 | 2023-02-01 | | 公開日 | 2023-12-27 | | 実験手法 | ELECTRON MICROSCOPY (1.98 Å) | | 主引用文献 | Efficient tagging of endogenous proteins in human cell lines for structural studies by single-particle cryo-EM.
Proc.Natl.Acad.Sci.USA, 120, 2023
|
|
8G14
 
 | |
8G13
 
 | |
8G12
 | |
8ZC5
 
 | | SARS-CoV-2 Omicron BA.4 spike trimer (6P) in complex with D1F6 Fab, focused refinement of RBD region | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of D1F6 Fab, Light chain of D1F6 Fab, ... | | 著者 | Liu, B, Gao, X, Li, Z, Chen, Q, He, J, Xiong, X. | | 登録日 | 2024-04-28 | | 公開日 | 2024-05-15 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (3.91 Å) | | 主引用文献 | An unconventional VH1-2 antibody tolerates escape mutations and shows an antigenic hotspot on SARS-CoV-2 spike.
Cell Rep, 43, 2024
|
|
8ZC3
 
 | | SARS-CoV-2 Omicron BA.4 spike trimer (6P) in complex with 3 D1F6 Fabs (1 RBD up) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of D1F6 Fab, ... | | 著者 | Liu, B, Gao, X, Li, Z, Chen, Q, He, J, Xiong, X. | | 登録日 | 2024-04-28 | | 公開日 | 2024-05-15 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (4.69 Å) | | 主引用文献 | An unconventional VH1-2 antibody tolerates escape mutations and shows an antigenic hotspot on SARS-CoV-2 spike.
Cell Rep, 43, 2024
|
|
8ZC0
 
 | | SARS-CoV-2 Omicron BA.2 spike trimer (6P) in complex with 3 D1F6 Fabs (2 RBD up) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of D1F6 Fab, ... | | 著者 | Liu, B, Gao, X, Li, Z, Chen, Q, He, J, Xiong, X. | | 登録日 | 2024-04-28 | | 公開日 | 2024-05-15 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (4.17 Å) | | 主引用文献 | An unconventional VH1-2 antibody tolerates escape mutations and shows an antigenic hotspot on SARS-CoV-2 spike.
Cell Rep, 43, 2024
|
|
8ZBZ
 
 | | SARS-CoV-2 Omicron BA.2 spike trimer (6P) in complex with 3 D1F6 Fabs (1 RBD up) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of D1F6 Fab, ... | | 著者 | Liu, B, Gao, X, Li, Z, Chen, Q, He, J, Xiong, X. | | 登録日 | 2024-04-28 | | 公開日 | 2024-05-15 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (4.71 Å) | | 主引用文献 | An unconventional VH1-2 antibody tolerates escape mutations and shows an antigenic hotspot on SARS-CoV-2 spike.
Cell Rep, 43, 2024
|
|
8ZC2
 
 | | SARS-CoV-2 Omicron BA.2 spike trimer (6P) in complex with D1F6 Fab, head-to-head aggregate | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of D1F6 Fab, ... | | 著者 | Liu, B, Gao, X, Li, Z, Chen, Q, He, J, Xiong, X. | | 登録日 | 2024-04-28 | | 公開日 | 2024-05-29 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (7.82 Å) | | 主引用文献 | An unconventional VH1-2 antibody tolerates escape mutations and shows an antigenic hotspot on SARS-CoV-2 spike.
Cell Rep, 43, 2024
|
|
8ZBY
 
 | | SARS-CoV-2 Omicron BA.1 spike trimer (x2-4P) in complex with 3 D1F6 Fabs (0 RBD up) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of D1F6 Fab, ... | | 著者 | Liu, B, Gao, X, Li, Z, Chen, Q, He, J, Xiong, X. | | 登録日 | 2024-04-28 | | 公開日 | 2024-05-15 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (3.67 Å) | | 主引用文献 | An unconventional VH1-2 antibody tolerates escape mutations and shows an antigenic hotspot on SARS-CoV-2 spike.
Cell Rep, 43, 2024
|
|
8ZC4
 
 | | SARS-CoV-2 Omicron BA.4 spike trimer (6P) in complex with 3 D1F6 Fabs (2 RBD up) | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Heavy chain of D1F6 Fab, ... | | 著者 | Liu, B, Gao, X, Li, Z, Chen, Q, He, J, Xiong, X. | | 登録日 | 2024-04-28 | | 公開日 | 2024-05-15 | | 最終更新日 | 2024-06-12 | | 実験手法 | ELECTRON MICROSCOPY (3.95 Å) | | 主引用文献 | An unconventional VH1-2 antibody tolerates escape mutations and shows an antigenic hotspot on SARS-CoV-2 spike.
Cell Rep, 43, 2024
|
|
7XIM
 | |
7WP6
 
 | | Cryo-EM structure of SARS-CoV-2 recombinant spike protein STFK in complex with three neutralizing antibodies | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 36H6 heavy chain, ... | | 著者 | Zheng, Q, Sun, H, Yuan, Q, Li, S, Xia, N. | | 登録日 | 2022-01-23 | | 公開日 | 2023-03-01 | | 最終更新日 | 2023-09-13 | | 実験手法 | ELECTRON MICROSCOPY (3.81 Å) | | 主引用文献 | Lineage-mosaic and mutation-patched spike proteins for broad-spectrum COVID-19 vaccine.
Cell Host Microbe, 30, 2022
|
|
7WP8
 
 | | Cryo-EM structure of SARS-CoV-2 recombinant spike protein STFK1628x in complex with three neutralizing antibodies | | 分子名称: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, 2B4 heavy chain, ... | | 著者 | Zheng, Q, Sun, H, Yuan, Q, Li, S, Xia, N. | | 登録日 | 2022-01-23 | | 公開日 | 2023-03-08 | | 最終更新日 | 2023-09-13 | | 実験手法 | ELECTRON MICROSCOPY (3.88 Å) | | 主引用文献 | Lineage-mosaic and mutation-patched spike proteins for broad-spectrum COVID-19 vaccine.
Cell Host Microbe, 30, 2022
|
|
