8CUN
 
 | |
8CWA
 
 | |
8CTO
 
 | |
8CUX
 
 | |
8CUV
 
 | |
8CUU
 
 | |
8CUT
 
 | |
8CUS
 
 | |
8CWZ
 
 | |
8CWS
 
 | |
8CWY
 
 | |
8CUW
 
 | |
8EOV
 
 | |
8EOZ
 
 | |
8EOX
 
 | |
6M6Z
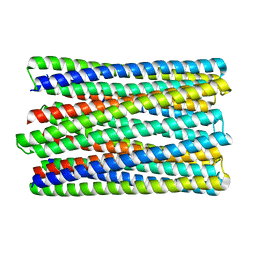 
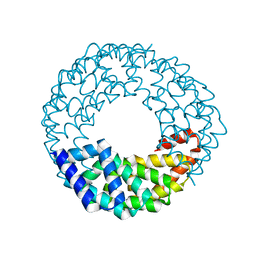
 | | A de novo designed transmembrane nanopore, TMH4C4 | | 分子名称: | TMH4C4 | | 著者 | Lu, P, Xu, C, Reggiano, G, Xu, Q, DiMaio, F, Baker, D. | | 登録日 | 2020-03-16 | | 公開日 | 2020-06-24 | | 最終更新日 | 2024-03-27 | | 実験手法 | ELECTRON MICROSCOPY (5.9 Å) | | 主引用文献 | Computational design of transmembrane pores.
Nature, 585, 2020
|
|
6MSP
 
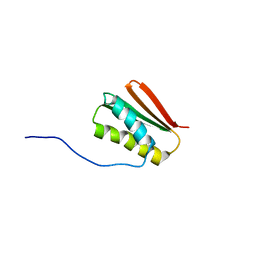 | | De novo Designed Protein Foldit3 | | 分子名称: | De novo Designed Protein Foldit3 | | 著者 | Liu, G, Ishida, Y, Swapna, G.V.T, Kleinfelter, S, Koepnick, B, Baker, D, Montelione, G.T. | | 登録日 | 2018-10-17 | | 公開日 | 2019-06-12 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | De novo protein design by citizen scientists.
Nature, 570, 2019
|
|
6MRS
 
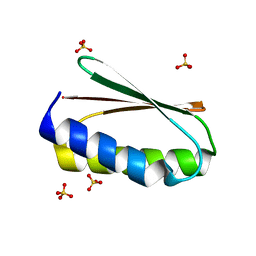 | |
6O0I
 
 | | NMR ensemble of computationally designed protein XAA | | 分子名称: | Design construct XAA | | 著者 | Wei, K.Y, Moschidi, D, Nerli, S, Sgourakis, N, Baker, D. | | 登録日 | 2019-02-16 | | 公開日 | 2020-04-22 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Computational design of closely related proteins that adopt two well-defined but structurally divergent folds.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6O0C
 
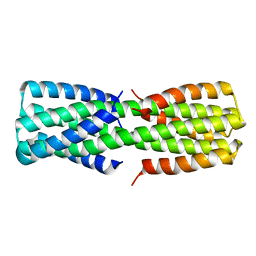 | | NMR ensemble of computationally designed protein XAA_GVDQ mutant M4L | | 分子名称: | Design construct XAA_GVDQ mutant M4L | | 著者 | Wei, K.Y, Moschidi, D, Nerli, S, Sgourakis, N, Baker, D. | | 登録日 | 2019-02-15 | | 公開日 | 2020-04-22 | | 最終更新日 | 2024-05-15 | | 実験手法 | SOLUTION NMR | | 主引用文献 | Computational design of closely related proteins that adopt two well-defined but structurally divergent folds.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
6VFJ
 
 | |
6VFH
 
 | |
6VFI
 
 | |
6V8E
 
 | |
6W3W
 
 | | An enumerative algorithm for de novo design of proteins with diverse pocket structures | | 分子名称: | DENOVO NTF2, NITRATE ION | | 著者 | Bera, A.K, Basanta, B, Dimaio, F, Sankaran, B, Baker, D. | | 登録日 | 2020-03-09 | | 公開日 | 2020-04-08 | | 最終更新日 | 2024-04-03 | | 実験手法 | X-RAY DIFFRACTION (1.55 Å) | | 主引用文献 | An enumerative algorithm for de novo design of proteins with diverse pocket structures.
Proc.Natl.Acad.Sci.USA, 117, 2020
|
|
