8DIG
 
 | |
8DIH
 
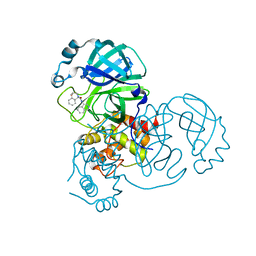
 | | Virtual screening for novel SARS-CoV-2 main protease non-covalent and covalent inhibitors | | 分子名称: | (1P,1'R)-1-(isoquinolin-4-yl)-2',3'-dihydrospiro[imidazolidine-4,1'-indene]-2,5-dione, 3C-like proteinase nsp5 | | 著者 | Singh, I, Shoichet, B.K. | | 登録日 | 2022-06-29 | | 公開日 | 2023-06-28 | | 最終更新日 | 2023-10-25 | | 実験手法 | X-RAY DIFFRACTION (2.12 Å) | | 主引用文献 | Large library docking for novel SARS-CoV-2 main protease non-covalent and covalent inhibitors.
Protein Sci., 32, 2023
|
|
8DIF
 
 | |
8DIE
 
 | |
8DII
 
 | |
8DID
 | |
7TWR
 
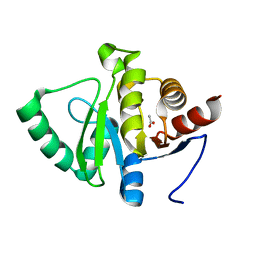 | |
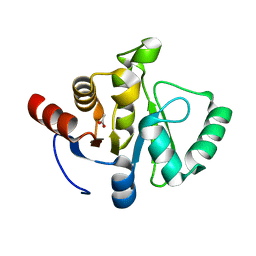
7TWP
 
 | |
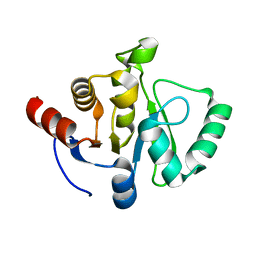
7TWG
 
 | |
7TWS
 
 | |
7TWF
 
 | |
7TWQ
 
 | |
7TWJ
 
 | |
7TWI
 
 | |
7TX5
 
 | |
7TWH
 
 | |
7TX3
 
 | |
7TWN
 
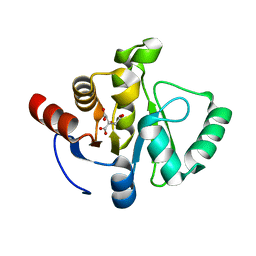 | |
7TWO
 
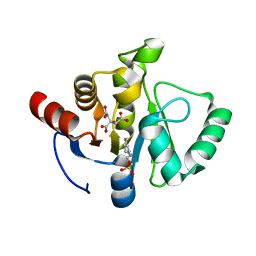 | |
7TX4
 
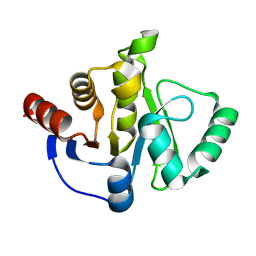 | |
7OU5
 
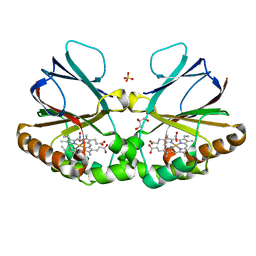 | | Crystal structure of dimeric chlorite dismutase from Cyanothece sp. PCC7425 in complex with nitrite | | 分子名称: | Chlorite Dismutase, GLYCEROL, NITRITE ION, ... | | 著者 | Schmidt, D, Mlynek, G, Djinovic-Carugo, K, Obinger, C. | | 登録日 | 2021-06-11 | | 公開日 | 2021-12-22 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.9 Å) | | 主引用文献 | Impact of the dynamics of the catalytic arginine on nitrite and chlorite binding by dimeric chlorite dismutase.
J.Inorg.Biochem., 227, 2021
|
|
7OU7
 
 | | Crystal structure of dimeric chlorite dismutase variant Q74V (CCld Q74V) from Cyanothece sp. PCC7425 in complex with nitrite | | 分子名称: | Chlorite dismutase, GLYCEROL, NITRITE ION, ... | | 著者 | Schmidt, D, Mlynek, G, Djinovic-Carugo, K, Obinger, C. | | 登録日 | 2021-06-11 | | 公開日 | 2021-12-22 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.63 Å) | | 主引用文献 | Impact of the dynamics of the catalytic arginine on nitrite and chlorite binding by dimeric chlorite dismutase.
J.Inorg.Biochem., 227, 2021
|
|
7OWI
 
 | | Crystal structure of dimeric chlorite dismutase variant R127A (CCld R127A) from Cyanothece sp. PCC7425 | | 分子名称: | (4S)-2-METHYL-2,4-PENTANEDIOL, Chlorite dismutase, DI(HYDROXYETHYL)ETHER, ... | | 著者 | Schmidt, D, Mlynek, G, Djinovic-Carugo, K, Obinger, C. | | 登録日 | 2021-06-18 | | 公開日 | 2021-12-22 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (1.7 Å) | | 主引用文献 | Impact of the dynamics of the catalytic arginine on nitrite and chlorite binding by dimeric chlorite dismutase.
J.Inorg.Biochem., 227, 2021
|
|
7OU9
 
 | | Crystal structure of dimeric chlorite dismutase variant Q74E (CCld Q74E) from Cyanothece sp. PCC7425 in complex with nitrite | | 分子名称: | Chlorite dismutase, GLYCEROL, NITRITE ION, ... | | 著者 | Schmidt, D, Mlynek, G, Djinovic-Carugo, K, Obinger, C. | | 登録日 | 2021-06-11 | | 公開日 | 2021-12-22 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (2.42 Å) | | 主引用文献 | Impact of the dynamics of the catalytic arginine on nitrite and chlorite binding by dimeric chlorite dismutase.
J.Inorg.Biochem., 227, 2021
|
|
7OUA
 
 | | Crystal structure of dimeric chlorite dismutase variant R127K (CCld R127K) from Cyanothece sp. PCC7425 | | 分子名称: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, Chlorite dismutase, DI(HYDROXYETHYL)ETHER, ... | | 著者 | Schmidt, D, Mlynek, G, Djinovic-Carugo, K, Obinger, C. | | 登録日 | 2021-06-11 | | 公開日 | 2021-12-22 | | 最終更新日 | 2024-01-31 | | 実験手法 | X-RAY DIFFRACTION (2.09 Å) | | 主引用文献 | Impact of the dynamics of the catalytic arginine on nitrite and chlorite binding by dimeric chlorite dismutase.
J.Inorg.Biochem., 227, 2021
|
|
