5X3O
 | |
5X3N
 
 | |
5X3M
 
 | |
5XPK
 
 | |
1Z7J
 
 | | Human transthyretin (also called prealbumin) complex with 3, 3',5,5'-tetraiodothyroacetic acid (t4ac) | | Descriptor: | 3,3',5,5'-TETRAIODOTHYROACETIC ACID, Transthyretin | | Authors: | Neumann, P, Wojtczak, A, Cody, V. | | Deposit date: | 2005-03-25 | | Release date: | 2005-06-07 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Ligand binding at the transthyretin dimer-dimer interface: structure of the transthyretin-T4Ac complex at 2.2 Angstrom resolution.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
3OC3
 
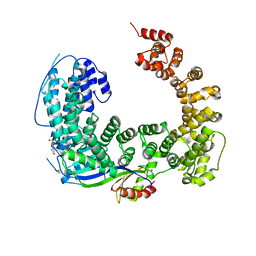 | | Crystal structure of the Mot1 N-terminal domain in complex with TBP | | Descriptor: | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID, HELICASE MOT1, TRANSCRIPTION INITIATION FACTOR TFIID (TFIID-1) | | Authors: | Wollmann, P, Cui, S, Viswanathan, R, Berninghausen, O, Wells, M.N, Moldt, M, Witte, G, Butryn, A, Wendler, P, Beckmann, R, Auble, D.T, Hopfner, K.-P. | | Deposit date: | 2010-08-09 | | Release date: | 2011-07-13 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structure and mechanism of the Swi2/Snf2 remodeller Mot1 in complex with its substrate TBP.
Nature, 475, 2011
|
|
3M63
 
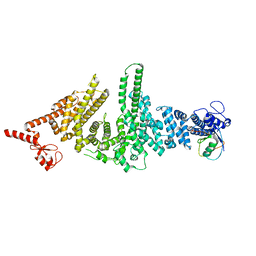 | |
3M62
 
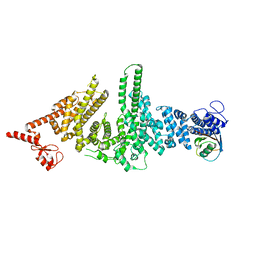 | |
3QQ7
 
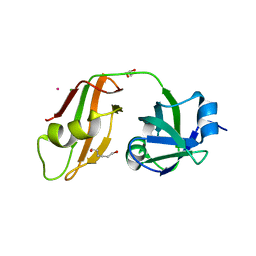 | |
3R3M
 | |
3QQ8
 
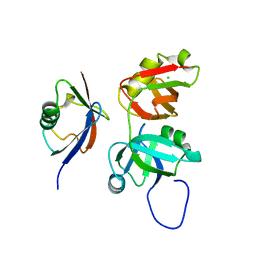 | |
3IPO
 
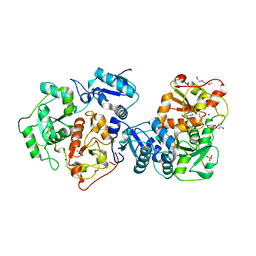 | | Crystal structure of YnjE | | Descriptor: | 2-(2-METHOXYETHOXY)ETHANOL, 2-{2-[2-(2-{2-[2-(2-ETHOXY-ETHOXY)-ETHOXY]-ETHOXY}-ETHOXY)-ETHOXY]-ETHOXY}-ETHANOL, GLYCEROL, ... | | Authors: | Haenzelmann, P, Kuper, J, Schindelin, H. | | Deposit date: | 2009-08-18 | | Release date: | 2009-12-08 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of YnjE from Escherichia coli, a sulfurtransferase with three rhodanese domains.
Protein Sci., 18, 2009
|
|
3IPP
 
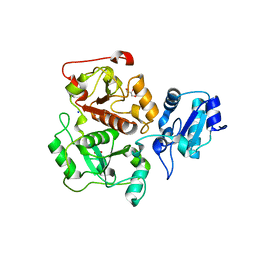 | | crystal structure of sulfur-free YnjE | | Descriptor: | GLYCEROL, PHOSPHATE ION, Putative thiosulfate sulfurtransferase ynjE, ... | | Authors: | Haenzelmann, P, Kuper, J, Schindelin, H. | | Deposit date: | 2009-08-18 | | Release date: | 2009-12-08 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Crystal structure of YnjE from Escherichia coli, a sulfurtransferase with three rhodanese domains.
Protein Sci., 18, 2009
|
|
2FB3
 
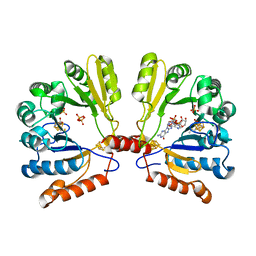 | | Structure of MoaA in complex with 5'-GTP | | Descriptor: | 5'-DEOXYADENOSINE, GUANOSINE-5'-TRIPHOSPHATE, IRON/SULFUR CLUSTER, ... | | Authors: | Haenzelmann, P, Schindelin, H. | | Deposit date: | 2005-12-08 | | Release date: | 2006-05-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.349 Å) | | Cite: | Binding of 5'-GTP to the C-terminal FeS cluster of the radical S-adenosylmethionine enzyme MoaA provides insights into its mechanism
Proc.Natl.Acad.Sci.USA, 103, 2006
|
|
2FB2
 
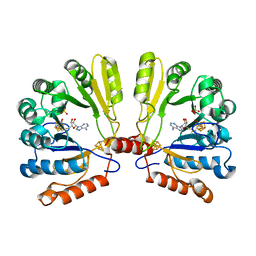 | | Structure of the MoaA Arg17/266/268/Ala triple mutant | | Descriptor: | IRON/SULFUR CLUSTER, Molybdenum cofactor biosynthesis protein A, S-ADENOSYLMETHIONINE, ... | | Authors: | Haenzelmann, P, Schindelin, H. | | Deposit date: | 2005-12-08 | | Release date: | 2006-05-09 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Binding of 5'-GTP to the C-terminal FeS cluster of the radical S-adenosylmethionine enzyme MoaA provides insights into its mechanism
Proc.Natl.Acad.Sci.USA, 103, 2006
|
|
4KR9
 
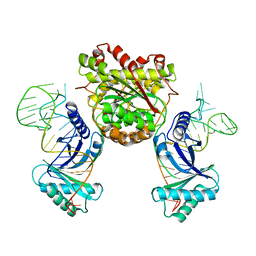 | |
5C19
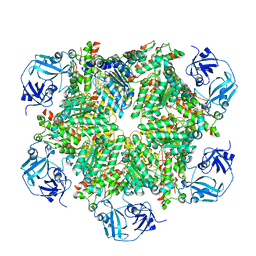 
 | | p97 variant 2 in the apo state | | Descriptor: | SULFATE ION, Transitional endoplasmic reticulum ATPase | | Authors: | Haenzelmann, P, Schindelin, H. | | Deposit date: | 2015-06-13 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (4.2 Å) | | Cite: | Structural Basis of ATP Hydrolysis and Intersubunit Signaling in the AAA+ ATPase p97.
Structure, 24, 2016
|
|
5C1B
 
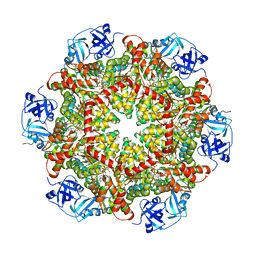 | |
8C4J
 
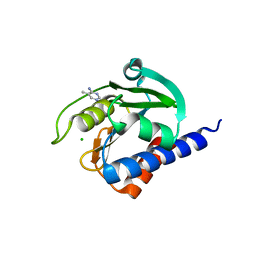 | | CdaA-compound 4 complex | | Descriptor: | 5-methylpyrimidin-4-amine, CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.83 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4N
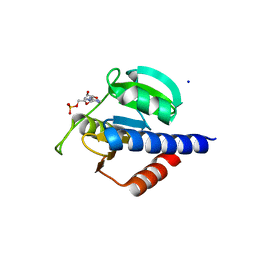 
 | | CdaA-AMP complex | | Descriptor: | ADENOSINE MONOPHOSPHATE, CHLORIDE ION, Diadenylate cyclase, ... | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4Q
 
 | | CdaA-Apo | | Descriptor: | CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4O
 
 | | CdaA-ATP complex | | Descriptor: | ADENOSINE-5'-TRIPHOSPHATE, CHLORIDE ION, Diadenylate cyclase, ... | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.97 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
8C4R
 
 | | CdaA-adenine complex | | Descriptor: | ADENINE, CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
5C1A
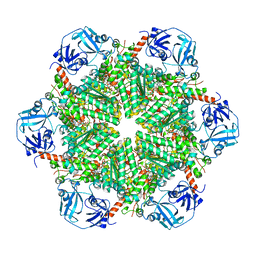 
 | | p97-N750D/R753D/M757D/Q760D in complex with ATP-gamma-S | | Descriptor: | MAGNESIUM ION, PHOSPHOTHIOPHOSPHORIC ACID-ADENYLATE ESTER, Transitional endoplasmic reticulum ATPase | | Authors: | Haenzelmann, P, Schindelin, H. | | Deposit date: | 2015-06-13 | | Release date: | 2016-01-13 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (3.8 Å) | | Cite: | Structural Basis of ATP Hydrolysis and Intersubunit Signaling in the AAA+ ATPase p97.
Structure, 24, 2016
|
|
8C4M
 
 | | CdaA-Adenosine complex | | Descriptor: | ADENOSINE, CHLORIDE ION, Diadenylate cyclase | | Authors: | Neumann, P, Ficner, R. | | Deposit date: | 2023-01-04 | | Release date: | 2023-06-07 | | Last modified: | 2024-06-19 | | Method: | X-RAY DIFFRACTION (1.51 Å) | | Cite: | Computer-aided design of a cyclic di-AMP synthesizing enzyme CdaA inhibitor.
Microlife, 4, 2023
|
|
