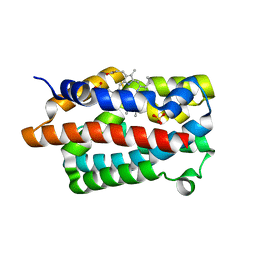
4GPH
 
 | |
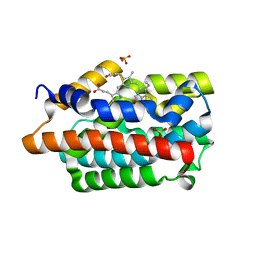
4GPC
 
 | |
3D6E
 
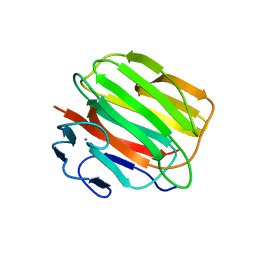 | | Crystal structure of the engineered 1,3-1,4-beta-glucanase protein from Bacillus licheniformis | | Descriptor: | Beta-glucanase, CALCIUM ION | | Authors: | Fita, I, Planas, A, Calisto, B.M, Addington, T. | | Deposit date: | 2008-05-19 | | Release date: | 2009-05-19 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Re-engineering specificity in 1,3-1,4-beta-glucanase to accept branched xyloglucan substrates
Proteins, 79, 2011
|
|
6MD6
 
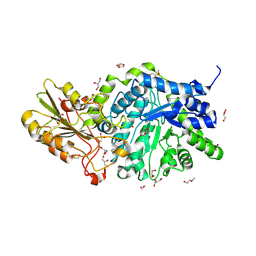 | | CRYSTAL STRUCTURE ANALYSIS OF PLANT EXOHYDROLASE IN COMPLEX WITH METHYL 2-THIO-BETA-SOPHOROSIDE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Beta-D-glucan exohydrolase isoenzyme ExoI, GLYCEROL, ... | | Authors: | Streltsov, V.A, Luang, S, Hrmova, M. | | Deposit date: | 2018-09-04 | | Release date: | 2019-05-29 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | Discovery of processive catalysis by an exo-hydrolase with a pocket-shaped active site.
Nat Commun, 10, 2019
|
|
6MI1
 
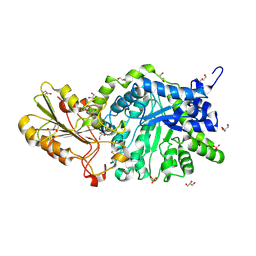 | | CRYSTAL STRUCTURE ANALYSIS OF THE VARIANT PLANT EXOHYDROLASE ARG158ALA-GLU161ALA IN COMPLEX WITH METHYL 6-THIO-BETA-GENTIOBIOSIDE | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, Beta-D-glucan exohydrolase isoenzyme ExoI, GLYCEROL, ... | | Authors: | Streltsov, V.A, Luang, S, Hrmova, M. | | Deposit date: | 2018-09-18 | | Release date: | 2019-05-29 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Discovery of processive catalysis by an exo-hydrolase with a pocket-shaped active site.
Nat Commun, 10, 2019
|
|
5V4O
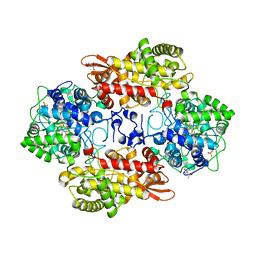 
 | |
5V53
 
 | |
9G98
 
 | | Joint neutron and x-ray structure of alginate lyase PsAlg7C soaked with pentamannuronic acid | | Descriptor: | 4-deoxy-alpha-L-erythro-hex-4-enopyranuronic acid-(1-4)-beta-D-mannopyranuronic acid, Alginate lyase, beta-D-mannopyranuronic acid-(1-4)-alpha-D-mannopyranuronic acid-(1-4)-beta-D-mannopyranuronic acid | | Authors: | Wilkens, C, Meilleur, F, Morth, J.P. | | Deposit date: | 2024-07-24 | | Release date: | 2025-07-30 | | Method: | NEUTRON DIFFRACTION (2.15 Å), X-RAY DIFFRACTION | | Cite: | Unraveling the molecular mechanism of polysaccharide lyases for efficient alginate degradation.
Nat Commun, 16, 2025
|
|
9G99
 
 | |
9FHX
 
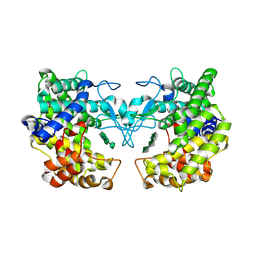 | |
9FHY
 
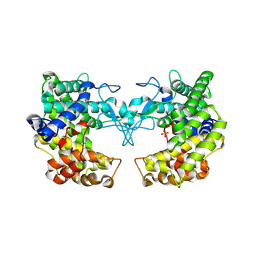 | |
9FHT
 
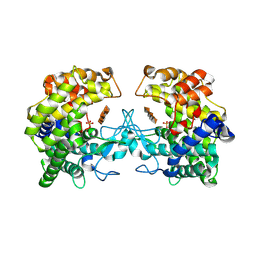 | |
9FHV
 
 | |
9FHU
 
 | |
9FI3
 
 | |
9FHZ
 
 | |
9FI1
 
 | |
9FI4
 
 | |
9FHW
 
 | |
9FI5
 
 | |
9FI0
 
 | |
9FI6
 
 | |
9FI2
 
 | |
7OOF
 
 | | Crystal structure of Paradendryphiella salina PL7A alginate lyase in complex with tri-mannuronic acid | | Descriptor: | Alginate lyase (PL7), TETRAETHYLENE GLYCOL, alpha-D-mannopyranuronic acid, ... | | Authors: | Fredslund, F, Welner, D.H, Wilkens, C. | | Deposit date: | 2021-05-27 | | Release date: | 2022-06-08 | | Last modified: | 2025-07-02 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Unraveling the molecular mechanism of polysaccharide lyases for efficient alginate degradation.
Nat Commun, 16, 2025
|
|
7ORY
 
 | |
