6XVM
 
 | |
6XVO
 
 | |
6XX4
 
 | |
6XX3
 
 | |
7PVT
 
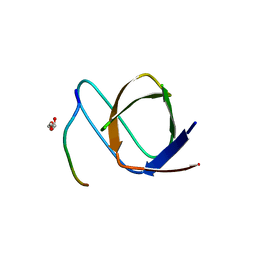 | |
6XX5
 
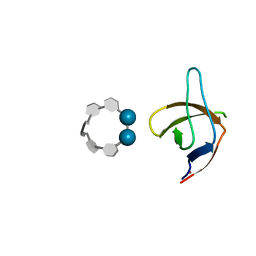 | |
6XX2
 
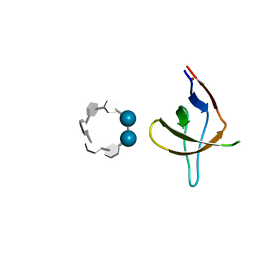 | |
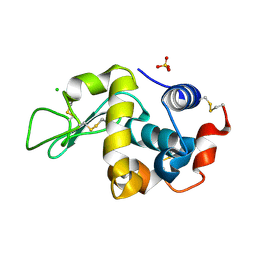
6F9Y
 
 | |
6F1L
 
 | |
6F1P
 
 | |
6F9Z
 
 | |
6F1M
 
 | |
6F1O
 
 | | Orthorhombic Lysozyme crystallized at 298 K and pH 4.5 | | Descriptor: | CHLORIDE ION, Lysozyme C, PHOSPHATE ION | | Authors: | Camara-Artigas, A. | | Deposit date: | 2017-11-22 | | Release date: | 2018-05-09 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (0.96 Å) | | Cite: | Orthorhombic lysozyme crystallization at acidic pH values driven by phosphate binding.
Acta Crystallogr D Struct Biol, 74, 2018
|
|
6F1R
 
 | |
6FA0
 
 | |
6F9X
 
 | |
