7V7K
 
 | | Crystal structure of Antibody 16A in complex with MUC1 Glycopeptide(GlycoST) | | Descriptor: | 16A fab heavy chain, 16A fab light chain, 2-acetamido-2-deoxy-beta-D-galactopyranose, ... | | Authors: | Niu, J, Xu, L, Meng, B, Han, Y.B, Yang, B. | | Deposit date: | 2021-08-21 | | Release date: | 2022-08-24 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Site-specific GalNAc modification on a MUC1 neoantigen epitope forms a basis for high-affinity antibody binding
To Be Published
|
|
7V8Q
 
 | | Crystal structure of antibody 14A in complex with MUC1 Glycopeptide(GlycoT) | | Descriptor: | 14A fab heavy chain, 14A fab light chain, 2-acetamido-2-deoxy-beta-D-galactopyranose, ... | | Authors: | Niu, J, Xu, L, Meng, B, Han, Y.B, Yang, B. | | Deposit date: | 2021-08-23 | | Release date: | 2022-08-31 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (3.2 Å) | | Cite: | Site-specific GalNAc modification on a MUC1 neoantigen epitope forms a basis for high-affinity antibody binding
To Be Published
|
|
7VAC
 
 | | Crystal structure of antibody 14A in complex with MUC1 glycopeptide(GlycoST) | | Descriptor: | 14A fab heavy chain, 14A fab light chain, 2-acetamido-2-deoxy-beta-D-galactopyranose, ... | | Authors: | Niu, J, Xu, L, Meng, B, Han, Y.B, Yang, B. | | Deposit date: | 2021-08-28 | | Release date: | 2022-08-31 | | Last modified: | 2024-10-16 | | Method: | X-RAY DIFFRACTION (3.5 Å) | | Cite: | Site-specific GalNAc modification on a MUC1 neoantigen epitope forms a basis for high-affinity antibody binding
To Be Published
|
|
7VAZ
 
 | | Crystal structure of antibody 14A in complex with MUC1 glycopeptide(GlycoS) | | Descriptor: | 14A fab heavy chain, 14A fab light chain, 2-acetamido-2-deoxy-beta-D-galactopyranose, ... | | Authors: | Niu, J, Xu, L, Meng, B, Han, Y.B, Yang, B. | | Deposit date: | 2021-08-30 | | Release date: | 2022-08-31 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.73 Å) | | Cite: | Site-specific GalNAc modification on a MUC1 neoantigen epitope forms a basis for high-affinity antibody binding
To Be Published
|
|
7V3Q
 
 | | Crystal structure of anti-MUC1 antibody 16A | | Descriptor: | Fab 16A Heavy Chain, Fab 16A Light Chain | | Authors: | Niu, J, Xu, L, Meng, B, Han, Y.B, Yang, B. | | Deposit date: | 2021-08-11 | | Release date: | 2022-08-17 | | Last modified: | 2024-10-09 | | Method: | X-RAY DIFFRACTION (2.98 Å) | | Cite: | Site-specific GalNAc modification on a MUC1 neoantigen epitope forms a basis for high-affinity antibody binding
To Be Published
|
|
7V4W
 
 | | Crystal structure of Antibody 16A in complex with MUC1 peptide | | Descriptor: | 16A Fab Heavy chain, 16A Fab Light chain, Mucin-1 subunit alpha | | Authors: | Niu, J, Xu, L, Meng, B, Han, Y.B, Yang, B. | | Deposit date: | 2021-08-15 | | Release date: | 2022-08-17 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Site-specific GalNAc modification on a MUC1 neoantigen epitope forms a basis for high-affinity antibody binding
To Be Published
|
|
1KAC
 
 | | KNOB DOMAIN FROM ADENOVIRUS SEROTYPE 12 IN COMPLEX WITH DOMAIN 1 OF ITS CELLULAR RECEPTOR CAR | | Descriptor: | PROTEIN (COXSACKIE VIRUS AND ADENOVIRUS RECEPTOR), PROTEIN (FIBER KNOB PROTEIN) | | Authors: | Bewley, M.C, Springer, K, Zhang, Y.B, Freimuth, P, Flanagan, J.M. | | Deposit date: | 1999-05-05 | | Release date: | 1999-11-24 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural analysis of the mechanism of adenovirus binding to its human cellular receptor, CAR.
Science, 286, 1999
|
|
1NOB
 
 | | KNOB DOMAIN FROM ADENOVIRUS SEROTYPE 12 | | Descriptor: | PROTEIN (FIBER KNOB PROTEIN) | | Authors: | Bewley, M.C, Springer, K, Zhang, Y.B, Freimuth, P, Flanagan, J.M. | | Deposit date: | 1999-05-05 | | Release date: | 1999-11-24 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural analysis of the mechanism of adenovirus binding to its human cellular receptor, CAR.
Science, 286, 1999
|
|
3SVL
 
 | | Structural basis of the improvement of ChrR - a multi-purpose enzyme | | Descriptor: | CALCIUM ION, FLAVIN MONONUCLEOTIDE, protein yieF | | Authors: | Poulain, S, Eswaramoorthy, S, Hienerwadel, R, Bremond, N, Sylvester, M.D, Zhang, Y.B, Van Der Lelie, D, Berthomieu, C, Matin, A.C. | | Deposit date: | 2011-07-12 | | Release date: | 2012-05-30 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal Structure of ChrR-A Quinone Reductase with the Capacity to Reduce Chromate.
Plos One, 7, 2012
|
|
5J14
 
 | | Crystal structure of endoglycoceramidase I from Rhodococ-cus equi in complex with GM3 | | Descriptor: | N-((E,2S,3R)-1,3-DIHYDROXYOCTADEC-4-EN-2-YL)STEARAMIDE, N-acetyl-alpha-neuraminic acid-(2-3)-beta-D-galactopyranose-(1-4)-beta-D-glucopyranose, Putative secreted endoglycosylceramidase, ... | | Authors: | Chen, L. | | Deposit date: | 2016-03-28 | | Release date: | 2016-04-27 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (1.915 Å) | | Cite: | Structural Insights into the Broad Substrate Specificity of a Novel Endoglycoceramidase I Belonging to a New Subfamily of GH5 Glycosidases
J. Biol. Chem., 292, 2017
|
|
5J7Z
 
 | | Crystal structure of endoglycoceramidase I from Rhodococ-cus equi in complex with GM1 | | Descriptor: | N-((E,2S,3R)-1,3-DIHYDROXYOCTADEC-4-EN-2-YL)STEARAMIDE, Putative secreted endoglycosylceramidase, SODIUM ION, ... | | Authors: | Chen, L. | | Deposit date: | 2016-04-07 | | Release date: | 2016-04-27 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Structural Insights into the Broad Substrate Specificity of a Novel Endoglycoceramidase I Belonging to a New Subfamily of GH5 Glycosidases.
J. Biol. Chem., 292, 2017
|
|
5CCU
 
 | | Crystal structure of endoglycoceramidase I from Rhodococ-cus equi | | Descriptor: | 1,2-ETHANEDIOL, Putative secreted endoglycosylceramidase, SODIUM ION | | Authors: | Chen, L. | | Deposit date: | 2015-07-02 | | Release date: | 2015-09-02 | | Last modified: | 2023-11-08 | | Method: | X-RAY DIFFRACTION (2.11 Å) | | Cite: | Structural Insights into the Broad Substrate Specificity of a Novel Endoglycoceramidase I Belonging to a New Subfamily of GH5 Glycosidases
J. Biol. Chem., 292, 2017
|
|
6Q1F
 
 | | Atomic structure of the Human Herpesvirus 6B Capsid and Capsid-Associated Tegument Complexes | | Descriptor: | Large structural phosphoprotein, Major capsid protein, Small capsomere-interacting protein, ... | | Authors: | Zhang, Y.B, Liu, W, Li, Z.H, Kumar, V, Alvarez-Cabrera, A.L, Leibovitch, E, Cui, Y.X, Mei, Y, Bi, G.Q, Jacobson, S, Zhou, Z.H. | | Deposit date: | 2019-08-03 | | Release date: | 2019-12-11 | | Last modified: | 2024-03-20 | | Method: | ELECTRON MICROSCOPY (9 Å) | | Cite: | Atomic structure of the human herpesvirus 6B capsid and capsid-associated tegument complexes.
Nat Commun, 10, 2019
|
|
5LG4
 
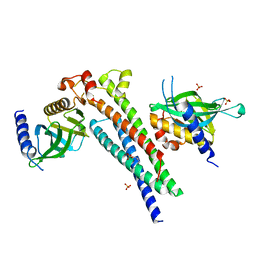 | |
6BMH
 
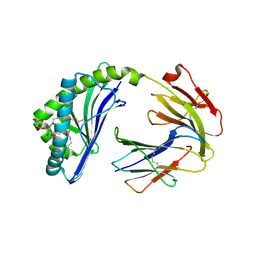 | | Crystal structure of MHC-I like protein | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, Antigen-presenting glycoprotein CD1d2, ... | | Authors: | Khandokar, Y.B, Le Nours, J, Rossjohn, J. | | Deposit date: | 2017-11-14 | | Release date: | 2018-01-31 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.298 Å) | | Cite: | Differing roles of CD1d2 and CD1d1 proteins in type I natural killer T cell development and function.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
6BMK
 
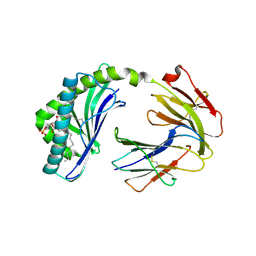 | | Crystal structure of MHC-I like protein | | Descriptor: | (2R)-1-(decanoyloxy)-3-(phosphonooxy)propan-2-yl octadecanoate, 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, ... | | Authors: | Khandokar, Y.B, Le Nours, J, Rossjohn, J. | | Deposit date: | 2017-11-15 | | Release date: | 2018-01-31 | | Last modified: | 2023-10-04 | | Method: | X-RAY DIFFRACTION (2.43 Å) | | Cite: | Differing roles of CD1d2 and CD1d1 proteins in type I natural killer T cell development and function.
Proc. Natl. Acad. Sci. U.S.A., 115, 2018
|
|
5M4Y
 
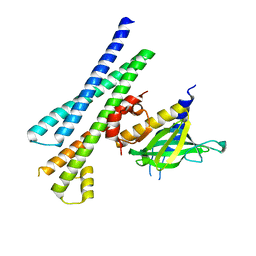 | |
2FR2
 
 | |
5Y3X
 
 | | Crystal structure of endo-1,4-beta-xylanase from Caldicellulosiruptor owensensis | | Descriptor: | Beta-xylanase | | Authors: | Liu, X, Sun, L.C, Zhang, Y.B, Liu, T.F, Xin, F.J. | | Deposit date: | 2017-07-31 | | Release date: | 2017-12-27 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structural Insights into the Thermophilic Adaption Mechanism of Endo-1,4-beta-Xylanase from Caldicellulosiruptor owensensis.
J. Agric. Food Chem., 66, 2018
|
|
1JEW
 
 | |
1P69
 
 | | STRUCTURAL BASIS FOR VARIATION IN ADENOVIRUS AFFINITY FOR THE CELLULAR RECEPTOR CAR (P417S MUTANT) | | Descriptor: | Coxsackievirus and adenovirus receptor, Fiber protein | | Authors: | Howitt, J, Bewley, M.C, Graziano, V, Flanagan, J.M, Freimuth, P. | | Deposit date: | 2003-04-29 | | Release date: | 2004-05-11 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Structural basis for variation in adenovirus affinity for the cellular coxsackievirus and adenovirus receptor.
J.Biol.Chem., 278, 2003
|
|
1P6A
 
 | | STRUCTURAL BASIS FOR VARIATION IN ADENOVIRUS AFFINITY FOR THE CELLULAR RECEPTOR CAR (S489Y MUTANT) | | Descriptor: | Coxsackievirus and adenovirus receptor, Fiber protein | | Authors: | Howitt, J, Bewley, M.C, Graziano, V, Flanagan, J.M, Freimuth, P. | | Deposit date: | 2003-04-29 | | Release date: | 2004-05-11 | | Last modified: | 2018-08-22 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Structural basis for variation in adenovirus affinity for the cellular coxsackievirus and adenovirus receptor.
J.Biol.Chem., 278, 2003
|
|
4MBU
 
 | |
4LUA
 
 | |
