1GBJ
 
 | | ALPHA-LYTIC PROTEASE WITH MET 190 REPLACED BY ALA | | Descriptor: | ALPHA-LYTIC PROTEASE, SULFATE ION | | Authors: | Mace, J.E, Agard, D.A. | | Deposit date: | 1995-09-06 | | Release date: | 1996-01-29 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Kinetic and structural characterization of mutations of glycine 216 in alpha-lytic protease: a new target for engineering substrate specificity.
J.Mol.Biol., 254, 1995
|
|
1GBC
 
 | |
1GBE
 
 | |
1GBH
 
 | |
1GBA
 
 | |
1GBM
 
 | |
2H5D
 | | 0.9A resolution crystal structure of alpha-lytic protease complexed with a transition state analogue, MeOSuc-Ala-Ala-Pro-Val boronic acid | | Descriptor: | ALPHA-LYTIC PROTEASE, GLYCEROL, MEOSUC-ALA-ALA-PRO-ALA BORONIC ACID INHIBITOR, ... | | Authors: | Fuhrmann, C.N, Agard, D.A. | | Deposit date: | 2006-05-25 | | Release date: | 2006-09-26 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (0.9 Å) | | Cite: | Subangstrom crystallography reveals that short ionic hydrogen bonds, and not a His-Asp low-barrier hydrogen bond, stabilize the transition state in serine protease catalysis
J.Am.Chem.Soc., 128, 2006
|
|
1BOQ
 | | PRO REGION C-TERMINUS: PROTEASE ACTIVE SITE INTERACTIONS ARE CRITICAL IN CATALYZING THE FOLDING OF ALPHA-LYTIC PROTEASE | | Descriptor: | PROTEIN (ALPHA-LYTIC PROTEASE), SULFATE ION | | Authors: | Peters, R.J, Shiau, A.K, Sohl, J.L, Anderson, D.E, Tang, G, Silen, J.L, Agard, D.A. | | Deposit date: | 1998-08-05 | | Release date: | 1998-08-12 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Pro region C-terminus:protease active site interactions are critical in catalyzing the folding of alpha-lytic protease.
Biochemistry, 37, 1998
|
|
1P01
 
 | |
1P10
 
 | |
1P09
 
 | |
1TAL
 
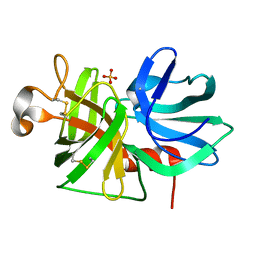
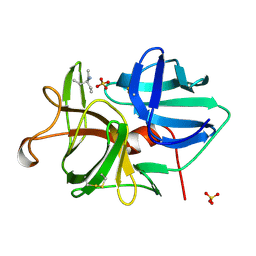 | | ALPHA-LYTIC PROTEASE AT 120 K (SINGLE STRUCTURE MODEL) | | Descriptor: | ALPHA-LYTIC PROTEASE, SULFATE ION, TRIS(HYDROXYETHYL)AMINOMETHANE | | Authors: | Rader, S.D, Agard, D.A. | | Deposit date: | 1996-10-30 | | Release date: | 1997-04-01 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | Conformational substates in enzyme mechanism: the 120 K structure of alpha-lytic protease at 1.5 A resolution.
Protein Sci., 6, 1997
|
|
3C7X
 
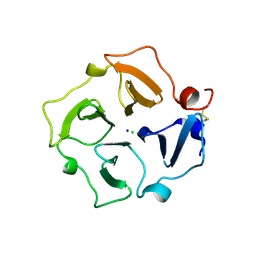 | | Hemopexin-like domain of matrix metalloproteinase 14 | | Descriptor: | CHLORIDE ION, Matrix metalloproteinase-14, SODIUM ION | | Authors: | Tochowicz, A, Itoh, Y, Maskos, K, Bode, W, Goettig, P. | | Deposit date: | 2008-02-08 | | Release date: | 2009-02-10 | | Last modified: | 2024-11-06 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | The dimer interface of the membrane type 1 matrix metalloproteinase hemopexin domain: crystal structure and biological functions
J.Biol.Chem., 286, 2011
|
|
6SZC
 
 | |
