2VQ5
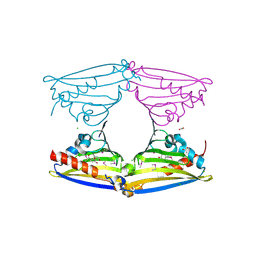 
 | | X-ray structure of Norcoclaurine synthase from Thalictrum flavum in complex with dopamine and hydroxybenzaldehyde | | Descriptor: | ACETATE ION, CHLORIDE ION, L-DOPAMINE, ... | | Authors: | Ilari, A, Franceschini, S, Bonamore, A, Boffi, A. | | Deposit date: | 2008-03-11 | | Release date: | 2008-08-05 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.09 Å) | | Cite: | Structural Basis of Enzymatic S-Norcoclaurine Biosynthesis.
J.Biol.Chem., 284, 2009
|
|
1IFV
 
 | |
2WQL
 
 | | CRYSTAL STRUCTURE OF THE MAJOR CARROT ALLERGEN DAU C 1 | | Descriptor: | CHLORIDE ION, DI(HYDROXYETHYL)ETHER, MAJOR ALLERGEN DAU C 1, ... | | Authors: | Markovic-Housley, Z, Basle, A, Padavattan, S, Hoffmann-Sommergruber, K, Schirmer, T. | | Deposit date: | 2009-08-24 | | Release date: | 2009-09-01 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Structure of the Major Carrot Allergen Dau C 1.
Acta Crystallogr.,Sect.D, 65, 2009
|
|
2I9Y
 
 | | Solution structure of Arabidopsis thaliana protein At1g70830, a member of the major latex protein family | | Descriptor: | major latex protein-like protein 28 or MLP-like protein 28 | | Authors: | Volkman, B.F, de la Cruz, N.B, Lytle, B.L, Peterson, F.C, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2006-09-06 | | Release date: | 2006-09-19 | | Last modified: | 2022-03-09 | | Method: | SOLUTION NMR | | Cite: | Structures of two Arabidopsis thaliana major latex proteins represent novel helix-grip folds.
Proteins, 76, 2009
|
|
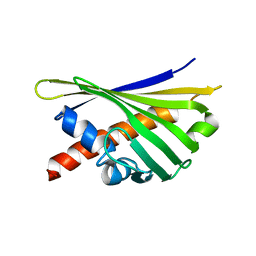
8QHH
 | |
8QHI
 
 | |
2Q3Q
 
 | | Ensemble refinement of the protein crystal structure of At1g24000 from Arabidopsis thaliana | | Descriptor: | Uncharacterized protein At1g24000 | | Authors: | Levin, E.J, Kondrashov, D.A, Wesenberg, G.E, Phillips Jr, G.N, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2007-05-30 | | Release date: | 2007-06-19 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Ensemble refinement of protein crystal structures: validation and application.
Structure, 15, 2007
|
|
1LLT
 
 | | BIRCH POLLEN ALLERGEN BET V 1 MUTANT E45S | | Descriptor: | POLLEN ALLERGEN BET V 1 | | Authors: | Spangfort, M.D, Mirza, O, Ipsen, H, Van Neerven, R.J, Gajhede, M, Larsen, J.N. | | Deposit date: | 2002-04-30 | | Release date: | 2003-10-28 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Dominating IgE-binding epitope of Bet v 1, the major allergen of birch pollen, characterized by X-ray crystallography and site-directed mutagenesis.
J.Immunol., 171, 2003
|
|
2QIM
 | | Crystal Structure of Pathogenesis-related Protein LlPR-10.2B from yellow lupine in complex with Cytokinin | | Descriptor: | (2E)-2-methyl-4-(9H-purin-6-ylamino)but-2-en-1-ol, CALCIUM ION, GLYCEROL, ... | | Authors: | Fernandes, H.C, Pasternak, O, Bujacz, G, Bujacz, A, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2007-07-05 | | Release date: | 2008-04-29 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.35 Å) | | Cite: | Lupinus luteus pathogenesis-related protein as a reservoir for cytokinin.
J.Mol.Biol., 378, 2008
|
|
2K7H
 | | NMR solution structure of soybean allergen Gly m 4 | | Descriptor: | Stress-induced protein SAM22 | | Authors: | Berkner, H, Neudecker, P, Mittag, D, Ballmer-Weber, B.K, Schweimer, K, Vieths, S, Roesch, P. | | Deposit date: | 2008-08-11 | | Release date: | 2009-05-05 | | Last modified: | 2022-03-16 | | Method: | SOLUTION NMR | | Cite: | Cross-reactivity of pollen and food allergens: soybean Gly m 4 is a member of the Bet v 1 superfamily and closely resembles yellow lupine proteins
Biosci.Rep., 29, 2009
|
|
1TXC
 
 | |
1TW0
 
 | |
1VJH
 
 | | Crystal structure of gene product of At1g24000 from Arabidopsis thaliana | | Descriptor: | Bet v I allergen family | | Authors: | Wesenberg, G.E, Smith, D.W, Phillips Jr, G.N, Johnson, K.A, Bingman, C.A, Center for Eukaryotic Structural Genomics (CESG) | | Deposit date: | 2004-02-20 | | Release date: | 2004-03-16 | | Last modified: | 2023-12-27 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | 1H, 15N and 13C resonance assignments of the putative Bet v 1 family protein At1g24000.1 from Arabidopsis thaliana.
J.Biomol.Nmr, 32, 2005
|
|
1XDF
 
 | | Crystal structure of pathogenesis-related protein LlPR-10.2A from yellow lupine | | Descriptor: | 4-(2-HYDROXYETHYL)-1-PIPERAZINE ETHANESULFONIC ACID, PR10.2A, SODIUM ION | | Authors: | Pasternak, O, Biesiadka, J, Dolot, R, Handschuh, L, Bujacz, G, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2004-09-06 | | Release date: | 2005-02-15 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Structure of a yellow lupin pathogenesis-related PR-10 protein belonging to a novel subclass.
Acta Crystallogr.,Sect.D, 61, 2005
|
|
4QIP
 
 | |
4REJ
 | |
4REI
 | |
6V8H
 | | Crystal structure of Ara h 8.0201 | | Descriptor: | Ara h 8 allergen isoform, SULFATE ION | | Authors: | Offermann, L.R, Pote, S, Hurlburt, B.K, McBride, J.K, Chruszcz, M. | | Deposit date: | 2019-12-11 | | Release date: | 2020-12-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.31 Å) | | Cite: | Crystal structure of Ara h 8.0201
To Be Published
|
|
6V8L
 | | Crystal structure of Ara h 8.0201 | | Descriptor: | Ara h 8 allergen isoform, SULFATE ION, icosanoic acid | | Authors: | Pote, S, Offermann, L.R, Hurlburt, B.K, McBride, J.K, Chruszcz, M. | | Deposit date: | 2019-12-11 | | Release date: | 2020-12-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of Ara h 8.0201
To Be Published
|
|
4REH
 | |
6V8J
 | | Crystal structure of Ara h 8.0201 | | Descriptor: | Ara h 8 allergen isoform, SODIUM ION, SULFATE ION | | Authors: | Offermann, L.R, Pote, S, Hurlburt, B.K, McBride, J.K, Chruszcz, M. | | Deposit date: | 2019-12-11 | | Release date: | 2020-12-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Crystal structure of Ara h 8.0201
To Be Published
|
|
6V8M
 | | Crystal structure of Ara h 8.0201 | | Descriptor: | 8-ANILINO-1-NAPHTHALENE SULFONATE, Ara h 8 allergen isoform, SULFATE ION | | Authors: | Pote, S, Offermann, L.R, Hurlburt, B.K, McBride, J.K, Chruszcz, M. | | Deposit date: | 2019-12-11 | | Release date: | 2020-12-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of Ara h 8.0201
To Be Published
|
|
6V8S
 | | Crystal structure of Ara h 8.0201 | | Descriptor: | 8-ANILINO-1-NAPHTHALENE SULFONATE, Ara h 8 allergen isoform, SULFATE ION | | Authors: | Pote, S, Offermann, L.R, Hurlburt, B.K, McBride, J.K, Chruszcz, M. | | Deposit date: | 2019-12-12 | | Release date: | 2020-12-16 | | Last modified: | 2023-10-11 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Crystal structure of Ara h 8.0201
To Be Published
|
|
4RYV
 
 | | Crystal structure of yellow lupin LLPR-10.1A protein in complex with trans-zeatin | | Descriptor: | (2E)-2-methyl-4-(9H-purin-6-ylamino)but-2-en-1-ol, Protein LLPR-10.1A, SULFATE ION | | Authors: | Dolot, R, Michalska, K, Sliwiak, J, Bujacz, G, Sikorski, M.M, Jaskolski, M. | | Deposit date: | 2014-12-17 | | Release date: | 2015-12-09 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (1.38 Å) | | Cite: | Crystallographic and CD probing of ligand-induced conformational changes in a plant PR-10 protein.
J.Struct.Biol., 193, 2016
|
|
5MMU
 | | NMR solution structure of the major apple allergen Mal d 1 | | Descriptor: | Major allergen Mal d 1 | | Authors: | Ahammer, L, Grutsch, S, Kamenik, A.S, Liedl, K.R, Tollinger, M. | | Deposit date: | 2016-12-12 | | Release date: | 2017-02-15 | | Last modified: | 2019-05-08 | | Method: | SOLUTION NMR | | Cite: | Structure of the Major Apple Allergen Mal d 1.
J. Agric. Food Chem., 65, 2017
|
|
