1JQI
 
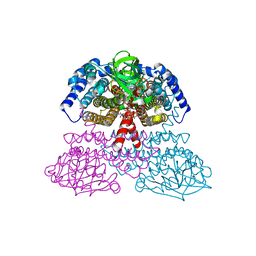 | | Crystal Structure of Rat Short Chain Acyl-CoA Dehydrogenase Complexed With Acetoacetyl-CoA | | Descriptor: | ACETOACETYL-COENZYME A, FLAVIN-ADENINE DINUCLEOTIDE, short chain acyl-CoA dehydrogenase | | Authors: | Battaile, K.P, Molin-Case, J, Paschke, R, Wang, M, Bennett, D, Vockley, J, Kim, J.-J.P. | | Deposit date: | 2001-08-07 | | Release date: | 2002-02-13 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.25 Å) | | Cite: | Crystal structure of rat short chain acyl-CoA dehydrogenase complexed with acetoacetyl-CoA: comparison with other acyl-CoA dehydrogenases.
J.Biol.Chem., 277, 2002
|
|
1JQJ
 
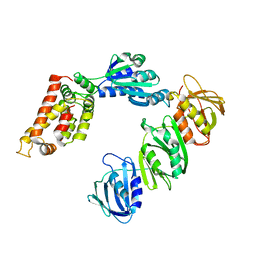 | | Mechanism of Processivity Clamp Opening by the Delta Subunit Wrench of the Clamp Loader Complex of E. coli DNA Polymerase III: Structure of the beta-delta complex | | Descriptor: | DNA polymerase III, beta chain, delta subunit | | Authors: | Jeruzalmi, D, Yurieva, O, Zhao, Y, Young, M, Stewart, J, Hingorani, M, O'Donnell, M, Kuriyan, J. | | Deposit date: | 2001-08-07 | | Release date: | 2001-11-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Mechanism of processivity clamp opening by the delta subunit wrench of the clamp loader complex of E. coli DNA polymerase III.
Cell(Cambridge,Mass.), 106, 2001
|
|
1JQL
 
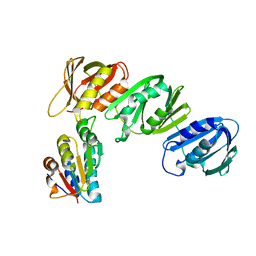 | | Mechanism of Processivity Clamp Opening by the Delta Subunit Wrench of the Clamp Loader Complex of E. coli DNA Polymerase III: Structure of beta-delta (1-140) | | Descriptor: | DNA Polymerase III, BETA CHAIN, DELTA SUBUNIT | | Authors: | Jeruzalmi, D, Yurieva, O, Zhao, Y, Young, M, Stewart, J, Hingorani, M, O'Donnell, M, Kuriyan, J. | | Deposit date: | 2001-08-07 | | Release date: | 2001-09-26 | | Last modified: | 2023-11-29 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Mechanism of processivity clamp opening by the delta subunit wrench of the clamp loader complex of E. coli DNA polymerase III.
Cell(Cambridge,Mass.), 106, 2001
|
|
1JQN
 
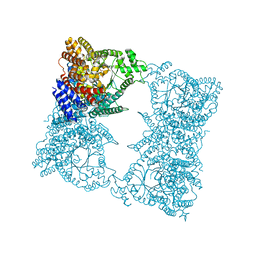 | |
1JQO
 
 | |
1JQP
 
 | | dipeptidyl peptidase I (cathepsin C), a tetrameric cysteine protease of the papain family | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, CHLORIDE ION, SULFATE ION, ... | | Authors: | Olsen, J.G, Kadziola, A, Lauritzen, C, Pedersen, J, Larsen, S, Dahl, S.W. | | Deposit date: | 2001-08-08 | | Release date: | 2002-10-18 | | Last modified: | 2020-07-29 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | Tetrameric dipeptidyl peptidase I directs substrate specificity by use of the residual pro-part domain
FEBS LETT., 506, 2001
|
|
1JQQ
 
 | | Crystal structure of Pex13p(301-386) SH3 domain | | Descriptor: | PEROXISOMAL MEMBRANE PROTEIN PAS20 | | Authors: | Douangamath, A, Mayans, O, Barnett, P, Distel, B, Wilmanns, M. | | Deposit date: | 2001-08-08 | | Release date: | 2002-12-06 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | Topography for Independent Binding of alpha-Helical and PPII-Helical Ligands to a Peroxisomal SH3 Domain
Mol.Cell, 10, 2002
|
|
1JQU
 
 | |
1JQV
 
 | | The K213E mutant of Lactococcus lactis Dihydroorotate dehydrogenase A | | Descriptor: | ACETIC ACID, Dihydroorotate dehydrogenase A, FLAVIN MONONUCLEOTIDE, ... | | Authors: | Norager, S, Arent, S, Bjornberg, O, Ottosen, M, Lo Leggio, L, Jensen, K.F, Larsen, S. | | Deposit date: | 2001-08-09 | | Release date: | 2003-09-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Lactococcus lactis dihydroorotate dehydrogenase A mutants reveal important facets of the enzymatic function
J.Biol.Chem., 278, 2003
|
|
1JQX
 
 | | The R57A mutant of Lactococcus lactis dihydroorotate dehydrogenase A | | Descriptor: | FLAVIN MONONUCLEOTIDE, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Norager, S, Arent, S, Bjornberg, O, Ottosen, M, Lo Leggio, L, Jensen, K.F, Larsen, S. | | Deposit date: | 2001-08-09 | | Release date: | 2003-09-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Lactococcus lactis dihydroorotate dehydrogenase A mutants reveal important facets of the enzymatic function
J.Biol.Chem., 278, 2003
|
|
1JQY
 
 | | HEAT-LABILE ENTEROTOXIN B-PENTAMER WITH LIGAND BMSC-0010 | | Descriptor: | (3-NITRO-5-(3-MORPHOLIN-4-YL-PROPYLAMINOCARBONYL)PHENYL)-GALACTOPYRANOSIDE, HEAT-LABILE ENTEROTOXIN B CHAIN | | Authors: | Merritt, E.A, Hol, W.G.J. | | Deposit date: | 2001-08-09 | | Release date: | 2002-05-08 | | Last modified: | 2017-10-04 | | Method: | X-RAY DIFFRACTION (2.14 Å) | | Cite: | Anchor-based design of improved cholera toxin and E. coli heat-labile enterotoxin receptor binding antagonists that display multiple binding modes.
Chem.Biol., 9, 2002
|
|
1JQZ
 
 | | Human Acidic Fibroblast Growth Factor. 141 Amino Acid Form with Amino Terminal His Tag. | | Descriptor: | FORMIC ACID, acidic fibroblast growth factor | | Authors: | Brych, S.R, Blaber, S.I, Logan, T.M, Blaber, M. | | Deposit date: | 2001-08-09 | | Release date: | 2001-12-19 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Structure and stability effects of mutations designed to increase the primary sequence symmetry within the core
region of a beta-trefoil.
Protein Sci., 10, 2001
|
|
1JR0
 
 | | CHOLERA TOXIN B-PENTAMER WITH LIGAND BMSC-0011 | | Descriptor: | (3-NITRO-5-(2-MORPHOLIN-4-YL-ETHYLAMINOCARBONYL)PHENYL)-GALACTOPYRANOSIDE, cholera toxin B subunit | | Authors: | Merritt, E.A, Hol, W.G.J. | | Deposit date: | 2001-08-09 | | Release date: | 2002-05-08 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.3 Å) | | Cite: | Anchor-based design of improved cholera toxin and E. coli heat-labile enterotoxin receptor binding antagonists that display multiple binding modes.
Chem.Biol., 9, 2002
|
|
1JR2
 
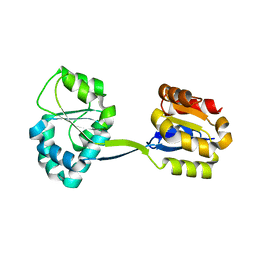 | | Structure of Uroporphyrinogen III Synthase | | Descriptor: | UROPORPHYRINOGEN-III SYNTHASE | | Authors: | Mathews, M.A, Schubert, H.L, Whitby, F.G, Alexander, K.J, Schadick, K, Bergonia, H.A, Phillips, J.D, Hill, C.P. | | Deposit date: | 2001-08-10 | | Release date: | 2001-11-07 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.84 Å) | | Cite: | Crystal structure of human uroporphyrinogen III synthase.
EMBO J., 20, 2001
|
|
1JR3
 
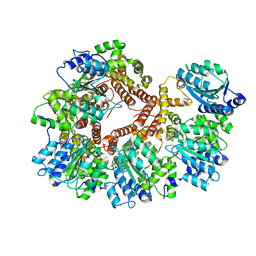 | | Crystal Structure of the Processivity Clamp Loader Gamma Complex of E. coli DNA Polymerase III | | Descriptor: | DNA polymerase III subunit gamma, DNA polymerase III, delta subunit, ... | | Authors: | Jeruzalmi, D, O'Donnell, M, Kuriyan, J. | | Deposit date: | 2001-08-10 | | Release date: | 2001-09-26 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structure of the processivity clamp loader gamma (gamma) complex of E. coli DNA polymerase III.
Cell(Cambridge,Mass.), 106, 2001
|
|
1JR4
 
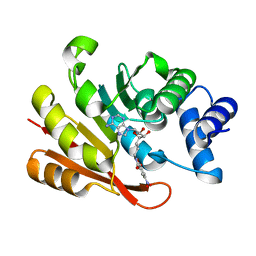 | | CATECHOL O-METHYLTRANSFERASE BISUBSTRATE-INHIBITOR COMPLEX | | Descriptor: | CATECHOL O-METHYLTRANSFERASE, SOLUBLE FORM, MAGNESIUM ION, ... | | Authors: | Ruf, A, Stihle, M. | | Deposit date: | 2001-08-10 | | Release date: | 2002-08-10 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.63 Å) | | Cite: | X-ray Crystal Structure of a Bisubstrate Inhibitor Bound to the Enzyme Catechol-O-methyltransferase: A Dramatic Effect of Inhibitor Preorganization on Binding Affinity.
Angew.Chem.Int.Ed.Engl., 40, 2001
|
|
1JR7
 
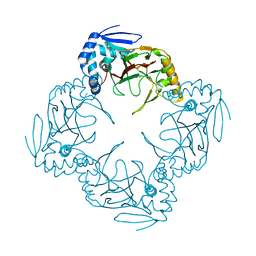 | |
1JR8
 
 | | Crystal Structure of Erv2p | | Descriptor: | Erv2 PROTEIN, mitochondrial, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Gross, E, Sevier, C.S, Vala, A, Kaiser, C.A, Fass, D. | | Deposit date: | 2001-08-13 | | Release date: | 2001-12-28 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.5 Å) | | Cite: | A new FAD-binding fold and intersubunit disulfide shuttle in the thiol oxidase Erv2p.
Nat.Struct.Biol., 9, 2002
|
|
1JR9
 
 | | Crystal Structure of manganese superoxide dismutases from Bacillus halodenitrificans | | Descriptor: | MANGANESE (II) ION, ZINC ION, manganese superoxide dismutase | | Authors: | Liao, J, Liu, M.Y, Chang, T, Li, M, LeGall, J, Gui, L.L, Zhang, J.P, Jiang, T, Liang, D.C, Chang, W.R. | | Deposit date: | 2001-08-13 | | Release date: | 2002-08-14 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Three-dimensional structure of manganese superoxide dismutase from Bacillus halodenitrificans, a component of the so-called "green protein".
J.Struct.Biol., 139, 2002
|
|
1JRA
 
 | | Crystal Structure of Erv2p | | Descriptor: | ERV2 PROTEIN, MITOCHONDRIAL, FLAVIN-ADENINE DINUCLEOTIDE | | Authors: | Gross, E, Sevier, C.S, Vala, A, Kaiser, C.A, Fass, D. | | Deposit date: | 2001-08-13 | | Release date: | 2001-12-28 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | A new FAD-binding fold and intersubunit disulfide shuttle in the thiol oxidase Erv2p.
Nat.Struct.Biol., 9, 2002
|
|
1JRB
 
 | | The P56A mutant of Lactococcus lactis dihydroorotate dehydrogenase A | | Descriptor: | FLAVIN MONONUCLEOTIDE, OROTIC ACID, dihydroorotate dehydrogenase A | | Authors: | Norager, S, Arent, S, Bjornberg, O, Ottosen, M, Lo Leggio, L, Jensen, K.F, Larsen, S. | | Deposit date: | 2001-08-13 | | Release date: | 2003-09-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.9 Å) | | Cite: | Lactococcus lactis dihydroorotate dehydrogenase A mutants reveal important facets of the enzymatic function
J.Biol.Chem., 278, 2003
|
|
1JRC
 
 | | The N67A mutant of Lactococcus lactis dihydroorotate dehydrogenase A | | Descriptor: | FLAVIN MONONUCLEOTIDE, OROTIC ACID, dihydroorotate dehydrogenase A | | Authors: | Norager, S, Arent, S, Bjornberg, O, Ottosen, M, Lo Leggio, L, Jensen, K.F, Larsen, S. | | Deposit date: | 2001-08-13 | | Release date: | 2003-09-09 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Lactococcus lactis dihydroorotate dehydrogenase A mutants reveal important facets of the enzymatic function
J.Biol.Chem., 278, 2003
|
|
1JRE
 
 | | DNA PROTECTION AND BINDING BY E. COLI DPS PROTEIN | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, CADMIUM ION, DNA PROTECTION DURING STARVATION PROTEIN | | Authors: | Luo, J, Liu, D, White, M.A, Fox, R.O. | | Deposit date: | 2001-08-13 | | Release date: | 2003-06-17 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.65 Å) | | Cite: | DNA Protection and Binding by E. Coli Dps Protein
To be Published
|
|
1JRG
 
 | | Crystal Structure of the R3 form of Pectate Lyase A, Erwinia chrysanthemi | | Descriptor: | Pectate lyase, SULFATE ION | | Authors: | Thomas, L.M, Doan, C, Oliver, R.L, Yoder, M.D. | | Deposit date: | 2001-08-13 | | Release date: | 2002-06-19 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structure of pectate lyase A: comparison to other isoforms.
Acta Crystallogr.,Sect.D, 58, 2002
|
|
1JRI
 
 | |
