1F6X
 
 | |
1F6Z
 
 | |
1F78
 | |
1F79
 
 | |
1F7E
 
 | | THE FIRST EGF-LIKE DOMAIN FROM HUMAN BLOOD COAGULATION FVII, NMR, 20 STRUCTURES | | Descriptor: | PROTEIN (Blood Coagulation Factor VII) | | Authors: | Kao, Y.-H, Lee, G.F, Wang, Y, Starovasnik, M.A, Kelley, R.F, Spellman, M.W, Lerner, L. | | Deposit date: | 1999-02-19 | | Release date: | 1999-06-16 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | The effect of O-fucosylation on the first EGF-like domain from human blood coagulation factor VII.
Biochemistry, 38, 1999
|
|
1F7F
 
 | |
1F7G
 
 | |
1F7H
 
 | |
1F7I
 
 | |
1F7M
 
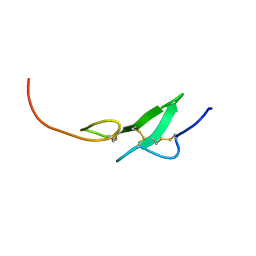 | | THE FIRST EGF-LIKE DOMAIN FROM HUMAN BLOOD COAGULATION FVII, NMR, MINIMIZED AVERAGE STRUCTURE | | Descriptor: | PROTEIN (Blood Coagulation Factor VII) | | Authors: | Kao, Y.-H, Lee, G.F, Wang, Y, Starovasnik, M.A, Kelley, R.F, Spellman, M.W, Lerner, L. | | Deposit date: | 1999-02-19 | | Release date: | 1999-06-16 | | Last modified: | 2024-11-13 | | Method: | SOLUTION NMR | | Cite: | The effect of O-fucosylation on the first EGF-like domain from human blood coagulation factor VII.
Biochemistry, 38, 1999
|
|
1F84
 
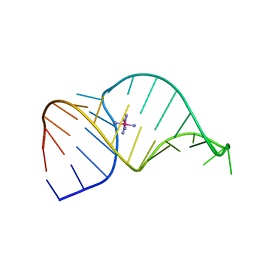
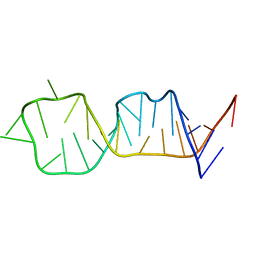 | | SOLUTION STRUCTURE OF HCV IRES RNA DOMAIN IIID | | Descriptor: | HCV-1B IRES RNA DOMAIN IIID | | Authors: | Lukavsky, P.J, Otto, G.A, Lancaster, A.M, Sarnow, P, Puglisi, J.D. | | Deposit date: | 2000-06-28 | | Release date: | 2000-11-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structures of two RNA domains essential for hepatitis C virus internal ribosome entry site function.
Nat.Struct.Biol., 7, 2000
|
|
1F85
 
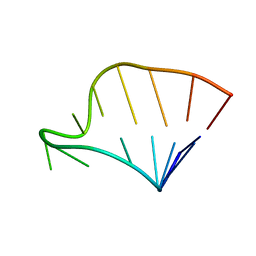 | | SOLUTION STRUCTURE OF HCV IRES RNA DOMAIN IIIE | | Descriptor: | HCV-1B IRES RNA DOMAIN IIIE | | Authors: | Lukavsky, P.J, Otto, G.A, Lancaster, A.M, Sarnow, P, Puglisi, J.D. | | Deposit date: | 2000-06-28 | | Release date: | 2000-11-17 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structures of two RNA domains essential for hepatitis C virus internal ribosome entry site function.
Nat.Struct.Biol., 7, 2000
|
|
1F8H
 
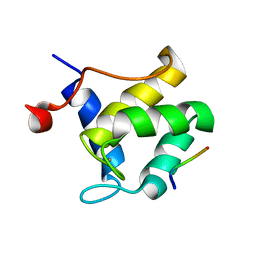 | | STRUCTURE OF THE SECOND EPS15 HOMOLOGY DOMAIN OF HUMAN EPS15 IN COMPLEX WITH PTGSSSTNPFR | | Descriptor: | CALCIUM ION, EPIDERMAL GROWTH FACTOR RECEPTOR SUBSTRATE 15, PTGSSSTNPFR | | Authors: | De Beer, T, Hoofnagle, A.N, Enmon, J.L, Bowers, R.C, Yamabhai, M, Kay, B.K, Overduin, M. | | Deposit date: | 2000-06-30 | | Release date: | 2000-11-01 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Molecular mechanism of NPF recognition by EH domains.
Nat.Struct.Biol., 7, 2000
|
|
1F8P
 | |
1F8Z
 
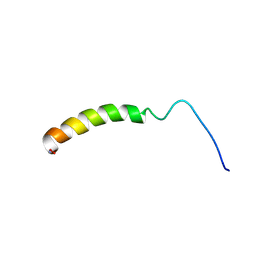
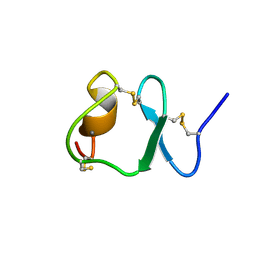 | | NMR STRUCTURE OF THE SIXTH LIGAND-BINDING MODULE OF THE LDL RECEPTOR | | Descriptor: | CALCIUM ION, LOW-DENSITY LIPOPROTEIN RECEPTOR | | Authors: | Clayton, D.J, Brereton, I.M, Kroon, P.A, Smith, R. | | Deposit date: | 2000-07-05 | | Release date: | 2000-10-18 | | Last modified: | 2024-10-30 | | Method: | SOLUTION NMR | | Cite: | Three-dimensional NMR structure of the sixth ligand-binding module of the human LDL receptor: comparison of two adjacent modules with different ligand binding specificities.
FEBS Lett., 479, 2000
|
|
1F95
 
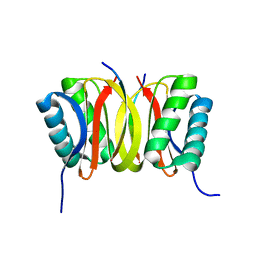 | | SOLUTION STRUCTURE OF DYNEIN LIGHT CHAIN 8 (DLC8) AND BIM PEPTIDE COMPLEX | | Descriptor: | BCL2-LIKE 11 (APOPTOSIS FACILITATOR), DYNEIN | | Authors: | Fan, J.-S, Zhang, Q, Tochio, H, Li, M, Zhang, M. | | Deposit date: | 2000-07-07 | | Release date: | 2001-02-28 | | Last modified: | 2024-05-01 | | Method: | SOLUTION NMR | | Cite: | Structural basis of diverse sequence-dependent target recognition by the 8 kDa dynein light chain.
J.Mol.Biol., 306, 2001
|
|
1F96
 
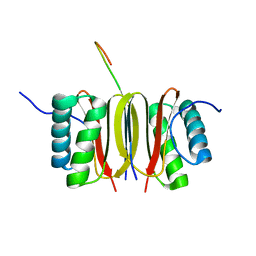 | | SOLUTION STRUCTURE OF DYNEIN LIGHT CHAIN 8 (DLC8) AND NNOS PEPTIDE COMPLEX | | Descriptor: | DYNEIN LIGHT CHAIN 8, PROTEIN (NNOS, NEURONAL NITRIC OXIDE SYNTHASE) | | Authors: | Fan, J.S, Zhang, Q, Tochio, H, Li, M, Zhang, M. | | Deposit date: | 2000-07-07 | | Release date: | 2001-02-28 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Structural basis of diverse sequence-dependent target recognition by the 8 kDa dynein light chain.
J.Mol.Biol., 306, 2001
|
|
1F9L
 
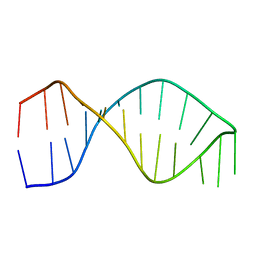 | |
1FA3
 
 | |
1FA4
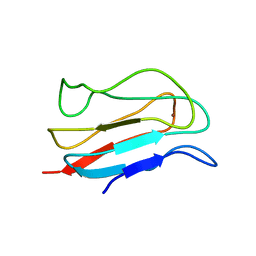 
 | | ELUCIDATION OF THE PARAMAGNETIC RELAXATION OF HETERONUCLEI AND PROTONS IN CU(II) PLASTOCYANIN FROM ANABAENA VARIABILIS | | Descriptor: | COPPER (II) ION, PLASTOCYANIN | | Authors: | Ma, L, Jorgensen, A.M, Sorensen, G.O, Ulstrup, J, Led, J.J. | | Deposit date: | 2000-07-12 | | Release date: | 2000-08-16 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | Elucidation of the Paramagnetic R1 Relaxation of Heteronuclei and Protons in Cu(II) Plastocyanin from Anabaena Variabilis
J.Am.Chem.Soc., 122, 2000
|
|
1FAD
 
 | | DEATH DOMAIN OF FAS-ASSOCIATED DEATH DOMAIN PROTEIN, RESIDUES 89-183 | | Descriptor: | PROTEIN (FADD PROTEIN) | | Authors: | Jeong, E.-J, Bang, S, Lee, T.H, Park, Y.-I, Sim, W.-S, Kim, K.-S. | | Deposit date: | 1999-03-23 | | Release date: | 1999-07-06 | | Last modified: | 2023-12-27 | | Method: | SOLUTION NMR | | Cite: | The solution structure of FADD death domain. Structural basis of death domain interactions of Fas and FADD.
J.Biol.Chem., 274, 1999
|
|
1FAF
 
 | | NMR STRUCTURE OF THE N-TERMINAL J DOMAIN OF MURINE POLYOMAVIRUS T ANTIGENS. | | Descriptor: | LARGE T ANTIGEN | | Authors: | Berjanskii, M.V, Riley, M.I, Xie, A, Semenchenko, V, Folk, W.R, Van Doren, S.R. | | Deposit date: | 2000-07-13 | | Release date: | 2000-11-22 | | Last modified: | 2024-05-22 | | Method: | SOLUTION NMR | | Cite: | NMR structure of the N-terminal J domain of murine polyomavirus T antigens. Implications for DnaJ-like domains and for mutations of T antigens.
J.Biol.Chem., 275, 2000
|
|
1FB9
 
 | |
1FC8
 
 | |
1FCL
 | |
