+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 1txx | ||||||
|---|---|---|---|---|---|---|---|
| タイトル | ACTIVE-SITE VARIANT OF E.COLI THIOREDOXIN | ||||||
 要素 要素 | PROTEIN (THIOREDOXIN) | ||||||
 キーワード キーワード | OXIDOREDUCTASE / REDOX / CXXC | ||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報DNA polymerase processivity factor activity / protein-disulfide reductase activity / cell redox homeostasis / cytoplasm / cytosol 類似検索 - 分子機能 | ||||||
| 生物種 |  | ||||||
| 手法 |  X線回折 / X線回折 /  分子置換 / 解像度: 2.2 Å 分子置換 / 解像度: 2.2 Å | ||||||
 データ登録者 データ登録者 | Schultz, L.W. / Chivers, P.T. / Raines, R.T. | ||||||
 引用 引用 |  ジャーナル: Acta Crystallogr.,Sect.D / 年: 1999 ジャーナル: Acta Crystallogr.,Sect.D / 年: 1999タイトル: The CXXC motif: crystal structure of an active-site variant of Escherichia coli thioredoxin. 著者: Schultz, L.W. / Chivers, P.T. / Raines, R.T. #1:  ジャーナル: Embo J. / 年: 1996 ジャーナル: Embo J. / 年: 1996タイトル: The Cxxc Motif: Imperatives for the Formation of Native Disulfide Bonds 著者: Chivers, P.T. / Labiossiere, M.C.A. / Raines, R.T. | ||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  1txx.cif.gz 1txx.cif.gz | 35.8 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb1txx.ent.gz pdb1txx.ent.gz | 23.3 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  1txx.json.gz 1txx.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/tx/1txx https://data.pdbj.org/pub/pdb/validation_reports/tx/1txx ftp://data.pdbj.org/pub/pdb/validation_reports/tx/1txx ftp://data.pdbj.org/pub/pdb/validation_reports/tx/1txx | HTTPS FTP |
|---|
-関連構造データ
| 関連構造データ | 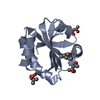 2trxS S: 精密化の開始モデル |
|---|---|
| 類似構造データ |
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| 単位格子 |
|
- 要素
要素
| #1: タンパク質 | 分子量: 11818.562 Da / 分子数: 1 / 変異: P33V, G34W / 由来タイプ: 組換発現 / 由来: (組換発現)    キーワード: OXIDOREDUCTASE キーワード: OXIDOREDUCTASE参照: UniProt: P0AA25, 酸化還元酵素; 含硫化合物に対し酸化酵素として働く |
|---|---|
| #2: 化合物 | ChemComp-CU / |
| #3: 水 | ChemComp-HOH / |
| Has protein modification | Y |
-実験情報
-実験
| 実験 | 手法:  X線回折 / 使用した結晶の数: 1 X線回折 / 使用した結晶の数: 1 |
|---|
- 試料調製
試料調製
| 結晶 | マシュー密度: 2.01 Å3/Da / 溶媒含有率: 37 % | ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 結晶化 | 手法: 蒸気拡散法, ハンギングドロップ法 / pH: 4.2 詳細: DROPS (6 ML) OF 0.050 M SODIUM SUCCINATE BUFFER, PH 4.2, CONTAINING CVWC THIOREDOXIN (10 MG/ML), METHYL-ETHER PEG2000 (10% W/V) AND CUPRIC ACETATE (1 MM) WERE SUSPENDED OVER 1.0 ML WELLS OF 0. ...詳細: DROPS (6 ML) OF 0.050 M SODIUM SUCCINATE BUFFER, PH 4.2, CONTAINING CVWC THIOREDOXIN (10 MG/ML), METHYL-ETHER PEG2000 (10% W/V) AND CUPRIC ACETATE (1 MM) WERE SUSPENDED OVER 1.0 ML WELLS OF 0.10 M SODIUM SUCCINATE BUFFER, PH 4.2, CONTAINING METHYL-ETHER PEG 2000 (20% W/V) AND CUPRIC ACETATE (2 MM)., VAPOR DIFFUSION, HANGING DROP | ||||||||||||||||||||||||||||||||||||||||
| 結晶 | *PLUS | ||||||||||||||||||||||||||||||||||||||||
| 結晶化 | *PLUS 温度: 293 K | ||||||||||||||||||||||||||||||||||||||||
| 溶液の組成 | *PLUS
|
-データ収集
| 回折 | 平均測定温度: 277 K |
|---|---|
| 放射光源 | 由来:  回転陽極 / タイプ: RIGAKU RU200 / 波長: 1.5418 回転陽極 / タイプ: RIGAKU RU200 / 波長: 1.5418 |
| 検出器 | タイプ: SIEMENS / 検出器: AREA DETECTOR / 日付: 1996年12月11日 / 詳細: MIRRORS |
| 放射 | プロトコル: SINGLE WAVELENGTH / 単色(M)・ラウエ(L): M / 散乱光タイプ: x-ray |
| 放射波長 | 波長: 1.5418 Å / 相対比: 1 |
| 反射 | 解像度: 2.2→30 Å / Num. obs: 5979 / % possible obs: 98 % / Observed criterion σ(I): 0.33 / 冗長度: 6.3 % / Rsym value: 0.083 / Net I/σ(I): 20 |
| 反射 シェル | 解像度: 2.2→2.3 Å / 冗長度: 3 % / Rsym value: 0.173 / % possible all: 95 |
| 反射 | *PLUS % possible obs: 98 % / Num. measured all: 37820 / Rmerge(I) obs: 0.083 |
| 反射 シェル | *PLUS % possible obs: 95 % / Rmerge(I) obs: 0.173 |
- 解析
解析
| ソフトウェア |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 構造決定の手法:  分子置換 分子置換開始モデル: 2TRX 解像度: 2.2→30 Å / σ(F): 0 / 立体化学のターゲット値: TNT PROTGEO /
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| 溶媒の処理 | Bsol: 275 Å2 / ksol: 0.82 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| 精密化ステップ | サイクル: LAST / 解像度: 2.2→30 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| 拘束条件 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||
| ソフトウェア | *PLUS 名称: TNT / バージョン: 5D / 分類: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||
| 精密化 | *PLUS 最高解像度: 2.2 Å / σ(F): 0 / Rfactor obs: 0.171 | ||||||||||||||||||||||||||||||||||||||||||||||||||
| 溶媒の処理 | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||
| 原子変位パラメータ | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||
| 拘束条件 | *PLUS
|
 ムービー
ムービー コントローラー
コントローラー













 PDBj
PDBj




