[English] 日本語
 Yorodumi
Yorodumi- SASDAL4: CRM1 RanGTP Snu1 (Exportin-1 + GTP-binding nuclear protein Ran + ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDAL4 |
|---|---|
 Sample Sample | CRM1 RanGTP Snu1
|
| Function / homology |  Function and homology information Function and homology informationEstrogen-dependent nuclear events downstream of ESR-membrane signaling / Cyclin A/B1/B2 associated events during G2/M transition / HuR (ELAVL1) binds and stabilizes mRNA / Heme signaling / cellular response to triglyceride / RNA import into nucleus / cellular response to salt / Transcriptional and post-translational regulation of MITF-M expression and activity / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Deactivation of the beta-catenin transactivating complex ...Estrogen-dependent nuclear events downstream of ESR-membrane signaling / Cyclin A/B1/B2 associated events during G2/M transition / HuR (ELAVL1) binds and stabilizes mRNA / Heme signaling / cellular response to triglyceride / RNA import into nucleus / cellular response to salt / Transcriptional and post-translational regulation of MITF-M expression and activity / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Deactivation of the beta-catenin transactivating complex / annulate lamellae / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / Resolution of Sister Chromatid Cohesion / regulation of proteasomal ubiquitin-dependent protein catabolic process / RHO GTPases Activate Formins / Separation of Sister Chromatids / RNA cap binding / RNA nuclear export complex / MAPK6/MAPK4 signaling / pre-miRNA export from nucleus / snRNA import into nucleus / nuclear export signal receptor activity / cellular response to mineralocorticoid stimulus / manchette / regulation of centrosome duplication / Regulation of cholesterol biosynthesis by SREBP (SREBF) / importin-alpha family protein binding / NLS-dependent protein nuclear import complex / regulation of protein export from nucleus / Rev-mediated nuclear export of HIV RNA / Nuclear import of Rev protein / protein localization to nucleolus / NEP/NS2 Interacts with the Cellular Export Machinery / tRNA processing in the nucleus / GTP metabolic process / Postmitotic nuclear pore complex (NPC) reformation / nuclear import signal receptor activity / protein-containing complex localization / nucleocytoplasmic transport / MicroRNA (miRNA) biogenesis / DNA metabolic process / dynein intermediate chain binding / protein complex oligomerization / regulation of protein catabolic process / mitotic sister chromatid segregation / viral process / spermatid development / ribosomal large subunit export from nucleus / nuclear pore / protein localization to nucleus / sperm flagellum / Cajal body / ribosomal subunit export from nucleus / mRNA export from nucleus / ribosomal small subunit export from nucleus / cytoskeleton organization / protein export from nucleus / mitotic spindle organization / male germ cell nucleus / hippocampus development / protein tetramerization / centriole / Transcriptional regulation by small RNAs / positive regulation of protein import into nucleus / recycling endosome / kinetochore / small GTPase binding / protein import into nucleus / melanosome / GDP binding / nuclear envelope / mitotic cell cycle / ribosome biogenesis / actin cytoskeleton organization / G protein activity / snRNP Assembly / midbody / nuclear membrane / DNA-binding transcription factor binding / Hydrolases; Acting on acid anhydrides; Acting on GTP to facilitate cellular and subcellular movement / cadherin binding / response to xenobiotic stimulus / ribonucleoprotein complex / protein heterodimerization activity / protein domain specific binding / cell division / GTPase activity / chromatin binding / GTP binding / chromatin / protein-containing complex binding / nucleolus / magnesium ion binding / negative regulation of transcription by RNA polymerase II / protein-containing complex / RNA binding / extracellular exosome / nucleoplasm / membrane Similarity search - Function |
| Biological species |   Homo sapiens (human) Homo sapiens (human) |
 Citation Citation |  Journal: Structure / Year: 2013 Journal: Structure / Year: 2013Title: Structural determinants and mechanism of mammalian CRM1 allostery. Authors: Nicole Dölker / Clement E Blanchet / Béla Voß / David Haselbach / Christian Kappel / Thomas Monecke / Dmitri I Svergun / Holger Stark / Ralf Ficner / Ulrich Zachariae / Helmut Grubmüller / Achim Dickmanns /  Abstract: Proteins carrying nuclear export signals cooperatively assemble with the export factor CRM1 and the effector protein RanGTP. In lower eukaryotes, this cooperativity is coupled to CRM1 conformational ...Proteins carrying nuclear export signals cooperatively assemble with the export factor CRM1 and the effector protein RanGTP. In lower eukaryotes, this cooperativity is coupled to CRM1 conformational changes; however, it is unknown if mammalian CRM1 maintains its compact conformation or shows similar structural flexibility. Here, combinations of small-angle X-ray solution scattering and electron microscopy experiments with molecular dynamics simulations reveal pronounced conformational flexibility in mammalian CRM1 and demonstrate that RanGTP binding induces association of its N- and C-terminal regions to form a toroid structure. The CRM1 toroid is stabilized mainly by local interactions between the terminal regions, rather than by global strain. The CRM1 acidic loop is key in transmitting the effect of this RanGTP-induced global conformational change to the NES-binding cleft by shifting its population to the open state, which displays enhanced cargo affinity. Cooperative CRM1 export complex assembly thus constitutes a highly dynamic process, encompassing an intricate interplay of global and local structural changes. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Data source
| SASBDB page |  SASDAL4 SASDAL4 |
|---|
-Related structure data
- External links
External links
| Related items in Molecule of the Month |
|---|
-Models
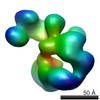
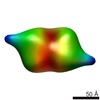
| Model #70 |  Type: dummy / Software: Dammif / Radius of dummy atoms: 1.90 A  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|---|
| Model #74 |  Type: dummy / Software: Monsa / Radius of dummy atoms: 1.90 A / Chi-square value: 1.5625  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
| Model #75 |  Type: dummy / Software: Monsa / Radius of dummy atoms: 1.90 A / Chi-square value: 1.550025  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
- Sample
Sample
 Sample Sample | Name: CRM1 RanGTP Snu1 / Sample MW: 188.65 kDa / Specimen concentration: 1.00-7.00 / Entity id: 62 / 63 / 64 |
|---|---|
| Buffer | Name: 50 mM Tris-HCL / pH: 7.5 / Composition: 150 mM NaCl, 1.0 mM DTT |
| Entity #62 | Type: protein / Description: Exportin-1 / Formula weight: 123.09 / Num. of mol.: 1 / Source: Mus musculus / References: UniProt: Q6P5F9 Sequence: MPAIMTMLAD HAARQLLDFS QKLDINLLDN VVNCLYHGEG AQQRMAQEVL THLKEHPDAW TRVDTILEFS QNMNTKYYGL QILENVIKTR WKILPRNQCE GIKKYVVGLI IKTSSDPTCV EKEKVYIGKL NMILVQILKQ EWPKHWPTFI SDIVGASRTS ESLCQNNMVI ...Sequence: MPAIMTMLAD HAARQLLDFS QKLDINLLDN VVNCLYHGEG AQQRMAQEVL THLKEHPDAW TRVDTILEFS QNMNTKYYGL QILENVIKTR WKILPRNQCE GIKKYVVGLI IKTSSDPTCV EKEKVYIGKL NMILVQILKQ EWPKHWPTFI SDIVGASRTS ESLCQNNMVI LKLLSEEVFD FSSGQITQVK AKHLKDSMCN EFSQIFQLCQ FVMENSQNAP LVHATLETLL RFLNWIPLGY IFETKLISTL IYKFLNVPMF RNVSLKCLTE IAGVSVSQYE EQFETLFTLT MMQLKQMLPL NTNIRLAYSN GKDDEQNFIQ NLSLFLCTFL KEHGQLLEKR LNLREALMEA LHYMLLVSEV EETEIFKICL EYWNHLAAEL YRESPFSTSA SPLLSGSQHF DIPPRRQLYL TVLSKVRLLM VSRMAKPEEV LVVENDQGEV VREFMKDTDS INLYKNMRET LVYLTHLDYV DTEIIMTKKL QNQVNGTEWS WKNLNTLCWA IGSISGAMHE EDEKRFLVTV IKDLLGLCEQ KRGKDNKAII ASNIMYIVGQ YPRFLRAHWK FLKTVVNKLF EFMHETHDGV QDMACDTFIK IAQKCRRHFV QVQVGEVMPF IDEILNNINT IICDLQPQQV HTFYEAVGYM IGAQTDQTVQ EHLIEKYMLL PNQVWDSIIQ QATKNVDILK DPETVKQLGS ILKTNVRACK AVGHPFVIQL GRIYLDMLNV YKCLSENISA AIQANGEMVT KQPLIRSMRT VKRETLKLIS GWVSRSNDPQ MVAENFVPPL LDAVLIDYQR NVPAAREPEV LSTMAIIVNK LGGHITAEIP QIFDAVFECT LNMINKDFEE YPEHRTNFFL LLQAVNSHCF PAFLAIPPAQ FKLVLDSIIW AFKHTMRNVA DTGLQILFTL LQNVAQEEAA AQSFYQTYFC DILQHIFSVV TDTSHTAGLT MHASILAYMF NLVEEGKIST PLNPGNPVNN QMFIQDYVAN LLKSAFPHLQ DAQVKLFVTG LFSLNQDIPA FKEHLRDFLV QIKEFAGEDT SDLFLEERET ALRQAQEEKH KLQMSVPGIL NPHEIPEEMC D |
| Entity #63 | Type: protein / Description: GTP-binding nuclear protein Ran / Formula weight: 24.42 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: P62826 Sequence: MAAQGEPQVQ FKLVLVGDGG TGKTTFVKRH LTGEFEKKYV ATLGVEVHPL VFHTNRGPIK FNVWDTAGQE KFGGLRDGYY IQAQCAIIMF DVTSRVTYKN VPNWHRDLVR VCENIPIVLC GNKVDIKDRK VKAKSIVFHR KKNLQYYDIS AKSNYNFEKP FLWLARKLIG ...Sequence: MAAQGEPQVQ FKLVLVGDGG TGKTTFVKRH LTGEFEKKYV ATLGVEVHPL VFHTNRGPIK FNVWDTAGQE KFGGLRDGYY IQAQCAIIMF DVTSRVTYKN VPNWHRDLVR VCENIPIVLC GNKVDIKDRK VKAKSIVFHR KKNLQYYDIS AKSNYNFEKP FLWLARKLIG DPNLEFVAMP ALAPPEVVMD PALAAQYEHD LEVAQTTALP DEDDDL |
| Entity #64 | Type: protein / Description: Snurportin-1 / Formula weight: 41.14 / Num. of mol.: 1 / Source: Homo sapiens / References: UniProt: O95149 Sequence: MEELSQALAS SFSVSQDLNS TAAPHPRLSQ YKSKYSSLEQ SERRRRLLEL QKSKRLDYVN HARRLAEDDW TGMESEEENK KDDEEMDIDT VKKLPKHYAN QLMLSEWLID VPSDLGQEWI VVVCPVGKRA LIVASRGSTS AYTKSGYCVN RFSSLLPGGN RRNSTAKDYT ...Sequence: MEELSQALAS SFSVSQDLNS TAAPHPRLSQ YKSKYSSLEQ SERRRRLLEL QKSKRLDYVN HARRLAEDDW TGMESEEENK KDDEEMDIDT VKKLPKHYAN QLMLSEWLID VPSDLGQEWI VVVCPVGKRA LIVASRGSTS AYTKSGYCVN RFSSLLPGGN RRNSTAKDYT ILDCIYNEVN QTYYVLDVMC WRGHPFYDCQ TDFRFYWMHS KLPEEEGLGE KTKLNPFKFV GLKNFPCTPE SLCDVLSMDF PFEVDGLLFY HKQTHYSPGS TPLVGWLRPY MVSDVLGVAV PAGPLTTKPD YAGHQLQQIM EHKKSQKEGM KEKLTHKASE NGHYELEHLS TPKLKGSSHS PDHPGCLMEN |
-Experimental information
| Beam | Instrument name:  DORIS III X33 DORIS III X33  / City: Hamburg / 国: Germany / City: Hamburg / 国: Germany  / Type of source: X-ray synchrotron / Type of source: X-ray synchrotron | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: MAR 345 Image Plate | ||||||||||||||||||||||||
| Scan |
| ||||||||||||||||||||||||
| Result | Type of curve: single_conc /
|
 Movie
Movie Controller
Controller