+Search query
-Structure paper
| Title | Structural determinants and mechanism of mammalian CRM1 allostery. |
|---|---|
| Journal, issue, pages | Structure, Vol. 21, Issue 8, Page 1350-1360, Year 2013 |
| Publish date | Aug 6, 2013 |
 Authors Authors | Nicole Dölker / Clement E Blanchet / Béla Voß / David Haselbach / Christian Kappel / Thomas Monecke / Dmitri I Svergun / Holger Stark / Ralf Ficner / Ulrich Zachariae / Helmut Grubmüller / Achim Dickmanns /  |
| PubMed Abstract | Proteins carrying nuclear export signals cooperatively assemble with the export factor CRM1 and the effector protein RanGTP. In lower eukaryotes, this cooperativity is coupled to CRM1 conformational ...Proteins carrying nuclear export signals cooperatively assemble with the export factor CRM1 and the effector protein RanGTP. In lower eukaryotes, this cooperativity is coupled to CRM1 conformational changes; however, it is unknown if mammalian CRM1 maintains its compact conformation or shows similar structural flexibility. Here, combinations of small-angle X-ray solution scattering and electron microscopy experiments with molecular dynamics simulations reveal pronounced conformational flexibility in mammalian CRM1 and demonstrate that RanGTP binding induces association of its N- and C-terminal regions to form a toroid structure. The CRM1 toroid is stabilized mainly by local interactions between the terminal regions, rather than by global strain. The CRM1 acidic loop is key in transmitting the effect of this RanGTP-induced global conformational change to the NES-binding cleft by shifting its population to the open state, which displays enhanced cargo affinity. Cooperative CRM1 export complex assembly thus constitutes a highly dynamic process, encompassing an intricate interplay of global and local structural changes. |
 External links External links |  Structure / Structure /  PubMed:23850451 PubMed:23850451 |
| Methods | SAS (X-ray synchrotron) / EM (single particle) |
| Resolution | 20.0 Å |
| Structure data |  SASDAJ4:  SASDAK4:  SASDAL4:  SASDAM4: 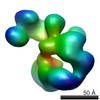 EMDB-2274:  EMDB-5564: |
| Source |
|
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers




 Homo sapiens (human)
Homo sapiens (human)