+ Open data
Open data
- Basic information
Basic information
| Entry | Database: SASBDB / ID: SASDCU7 |
|---|---|
 Sample Sample | TRIP18(7.5R)SN-f5
|
| Biological species | synthetic construct (others) |
 Citation Citation |  Journal: Nat Biotechnol / Year: 2017 Journal: Nat Biotechnol / Year: 2017Title: Design of coiled-coil protein-origami cages that self-assemble in vitro and in vivo. Authors: Ajasja Ljubetič / Fabio Lapenta / Helena Gradišar / Igor Drobnak / Jana Aupič / Žiga Strmšek / Duško Lainšček / Iva Hafner-Bratkovič / Andreja Majerle / Nuša Krivec / Mojca ...Authors: Ajasja Ljubetič / Fabio Lapenta / Helena Gradišar / Igor Drobnak / Jana Aupič / Žiga Strmšek / Duško Lainšček / Iva Hafner-Bratkovič / Andreja Majerle / Nuša Krivec / Mojca Benčina / Tomaž Pisanski / Tanja Ćirković Veličković / Adam Round / José María Carazo / Roberto Melero / Roman Jerala /      Abstract: Polypeptides and polynucleotides are natural programmable biopolymers that can self-assemble into complex tertiary structures. We describe a system analogous to designed DNA nanostructures in which ...Polypeptides and polynucleotides are natural programmable biopolymers that can self-assemble into complex tertiary structures. We describe a system analogous to designed DNA nanostructures in which protein coiled-coil (CC) dimers serve as building blocks for modular de novo design of polyhedral protein cages that efficiently self-assemble in vitro and in vivo. We produced and characterized >20 single-chain protein cages in three shapes-tetrahedron, four-sided pyramid, and triangular prism-with the largest containing >700 amino-acid residues and measuring 11 nm in diameter. Their stability and folding kinetics were similar to those of natural proteins. Solution small-angle X-ray scattering (SAXS), electron microscopy (EM), and biophysical analysis confirmed agreement of the expressed structures with the designs. We also demonstrated self-assembly of a tetrahedral structure in bacteria, mammalian cells, and mice without evidence of inflammation. A semi-automated computational design platform and a toolbox of CC building modules are provided to enable the design of protein cages in any polyhedral shape. |
 Contact author Contact author |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
-Models
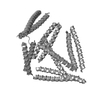
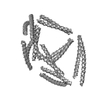
| Model #1584 |  Type: atomic / Software: (9.16) / Radius of dummy atoms: 1.90 A Comment: The model contributes 76% to the observed scattering curve. Chi-square value: 0.96  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
|---|---|
| Model #1585 | 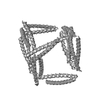 Type: atomic / Software: (9.16) / Radius of dummy atoms: 1.90 A Comment: The model contributes 24% to the observed scattering curve. Chi-square value: 0.96  Search similar-shape structures of this assembly by Omokage search (details) Search similar-shape structures of this assembly by Omokage search (details) |
- Sample
Sample
 Sample Sample | Name: TRIP18(7.5R)SN-f5 / Specimen concentration: 0.40-4.10 |
|---|---|
| Buffer | Name: 20 mM Tris 150 mM NaCl 10% glycerol / pH: 7.5 |
| Entity #807 | Name: TRIP18SN / Type: protein / Description: TRIP18(7.5R)SN-f5 / Formula weight: 81.866 / Num. of mol.: 1 / Source: synthetic construct Sequence: MSPEDENSQL EEKISQLKQK NSELKEEIQQ LEYGSGPGLE QIEERLEQIE ERLQAKEWEK AQLREELQAL REKLAQLGSG PGSPEDEIRQ LEQENSQLER ENQRLEQEIY QLERGSGPGS PEDKNSELKE EIQQLEEENQ QLEEKISELK YGSGPGSRMK QLEDKVEELL ...Sequence: MSPEDENSQL EEKISQLKQK NSELKEEIQQ LEYGSGPGLE QIEERLEQIE ERLQAKEWEK AQLREELQAL REKLAQLGSG PGSPEDEIRQ LEQENSQLER ENQRLEQEIY QLERGSGPGS PEDKNSELKE EIQQLEEENQ QLEEKISELK YGSGPGSRMK QLEDKVEELL SKNYHLENEV ERLKKLVGSG PGLEEELKQL EEELQAIEEQ LAQLQWKAQA RKEKLAQLKE KLGSGPGSPE DENQSLEQKN SQLKQEISQL EQEIQQLEYG SGPGLEQIEE RLEQIEERLQ AKEWEKAQLR EELQALREKL AQLGSGPGSR MKQLEDKVEE LLSKNYHLEN EVERLKKLVG SGPGSPEDEI QQLEEEISQL EQKNSELKEK NQELKYGSGP GSPEDENQSL EQEISQLEQE IQQLEQKNSE LKYGSGPGSP EDKNSQLKEE NSQLEEKIEQ LKEKIQELKY GSGPGSPEDK IEELKEKNSQ LKEKNEELKQ KIYELKEGSG PGDIEQELER AKESIRRLEQ EVNQERSRMQ YLQTLLEKGS GPGSPEDKNE QLKEKISELK EKIEQLKEEN QSLEYGSGPG LEEELKQLEE ELQAIEEQLA QLQWKAQARK EKLAQLKEKL GSGPGSPEDK ISQLKEKIQQ LKQENQQLEE ENSQLEYGSG PGDIEQELER AKESIRRLEQ EVNQERSRMQ YLQTLLEKLE HHHHHHHH |
-Experimental information
| Beam | Instrument name: PETRA III P12 / City: Hamburg / 国: Germany  / Shape: Point / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 2 mm / Shape: Point / Type of source: X-ray synchrotron / Wavelength: 0.124 Å / Dist. spec. to detc.: 2 mm | ||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Detector | Name: Pilatus 1M-W / Pixsize x: 0.172 mm | ||||||||||||||||||||||||||||||||||||||||||
| Scan |
| ||||||||||||||||||||||||||||||||||||||||||
| Distance distribution function P(R) |
| ||||||||||||||||||||||||||||||||||||||||||
| Result |
|
 Movie
Movie Controller
Controller


 SASDCU7
SASDCU7