[English] 日本語
 Yorodumi
Yorodumi- PDB-7n5s: ZBTB7A Zinc Finger Domain Bound to -200 Site of Fetal Globin Prom... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7n5s | ||||||
|---|---|---|---|---|---|---|---|
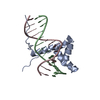
| Title | ZBTB7A Zinc Finger Domain Bound to -200 Site of Fetal Globin Promoter (Oligo 6) | ||||||
 Components Components |
| ||||||
 Keywords Keywords | DNA BINDING PROTEIN/DNA / Z=zinc-finger domain / gene expression / DNA BINDING PROTEIN-DNA complex | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of transcription regulatory region DNA binding / DNA-dependent protein kinase complex / negative regulation of androgen receptor signaling pathway / : / double-strand break repair via classical nonhomologous end joining / regulation of glycolytic process / nuclear androgen receptor binding / erythrocyte maturation / histone acetyltransferase binding / regulation of alternative mRNA splicing, via spliceosome ...regulation of transcription regulatory region DNA binding / DNA-dependent protein kinase complex / negative regulation of androgen receptor signaling pathway / : / double-strand break repair via classical nonhomologous end joining / regulation of glycolytic process / nuclear androgen receptor binding / erythrocyte maturation / histone acetyltransferase binding / regulation of alternative mRNA splicing, via spliceosome / SMAD binding / fat cell differentiation / negative regulation of Notch signaling pathway / protein localization to nucleus / B cell differentiation / transcription corepressor binding / negative regulation of transforming growth factor beta receptor signaling pathway / : / DNA-binding transcription repressor activity, RNA polymerase II-specific / sequence-specific double-stranded DNA binding / site of double-strand break / chromatin organization / regulation of apoptotic process / DNA-binding transcription factor binding / DNA-binding transcription factor activity, RNA polymerase II-specific / RNA polymerase II cis-regulatory region sequence-specific DNA binding / chromatin remodeling / DNA-binding transcription factor activity / negative regulation of DNA-templated transcription / DNA-templated transcription / regulation of transcription by RNA polymerase II / negative regulation of transcription by RNA polymerase II / DNA binding / zinc ion binding / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.86 Å MOLECULAR REPLACEMENT / Resolution: 2.86 Å | ||||||
 Authors Authors | Horton, J.R. / Ren, R. / Cheng, X. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Cell Rep / Year: 2021 Journal: Cell Rep / Year: 2021Title: Structural basis for human ZBTB7A action at the fetal globin promoter. Authors: Yang, Y. / Ren, R. / Ly, L.C. / Horton, J.R. / Li, F. / Quinlan, K.G.R. / Crossley, M. / Shi, Y. / Cheng, X. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7n5s.cif.gz 7n5s.cif.gz | 110.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7n5s.ent.gz pdb7n5s.ent.gz | 67.5 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7n5s.json.gz 7n5s.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/n5/7n5s https://data.pdbj.org/pub/pdb/validation_reports/n5/7n5s ftp://data.pdbj.org/pub/pdb/validation_reports/n5/7n5s ftp://data.pdbj.org/pub/pdb/validation_reports/n5/7n5s | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  7eyiC  7n5tC 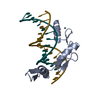 7n5uS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 16638.525 Da / Num. of mol.: 1 / Fragment: zinc finger domain (UNP residues 369-500) Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: ZBTB7A, FBI1, LRF, ZBTB7, ZNF857A / Production host: Homo sapiens (human) / Gene: ZBTB7A, FBI1, LRF, ZBTB7, ZNF857A / Production host:  | ||||
|---|---|---|---|---|---|
| #2: DNA chain | Mass: 4676.021 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) Homo sapiens (human) | ||||
| #3: DNA chain | Mass: 4506.912 Da / Num. of mol.: 1 / Source method: obtained synthetically / Source: (synth.)  Homo sapiens (human) Homo sapiens (human) | ||||
| #4: Chemical | ChemComp-ZN / #5: Water | ChemComp-HOH / | Has ligand of interest | N | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.46 Å3/Da / Density % sol: 49.93 % |
|---|---|
| Crystal grow | Temperature: 289 K / Method: vapor diffusion, sitting drop / pH: 8.5 Details: 0.2 M trimethylamine N-oxide dihydrate, 0.1 M Tris, pH 8.5, 20% w/v PEG2000 MME |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 1 Å / Beamline: 22-ID / Wavelength: 1 Å |
| Detector | Type: DECTRIS EIGER X 16M / Detector: PIXEL / Date: Dec 14, 2020 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.85→36.32 Å / Num. obs: 10160 / % possible obs: 92.6 % / Redundancy: 10.3 % / Biso Wilson estimate: 107.27 Å2 / Rmerge(I) obs: 0.097 / Rpim(I) all: 0.03 / Net I/σ(I): 18.8 |
| Reflection shell | Resolution: 2.85→2.95 Å / Redundancy: 4.1 % / Rmerge(I) obs: 1.67 / Mean I/σ(I) obs: 1.4 / Num. unique obs: 628 / CC1/2: 0.61 / Rpim(I) all: 0.701 / % possible all: 66 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB entry 7N5U Resolution: 2.86→36.32 Å / SU ML: 0.4711 / Cross valid method: FREE R-VALUE / σ(F): 1.36 / Phase error: 40.628 Stereochemistry target values: GeoStd + Monomer Library + CDL v1.2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 113.55 Å2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.86→36.32 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Refine-ID: X-RAY DIFFRACTION
|
 Movie
Movie Controller
Controller









 PDBj
PDBj













































