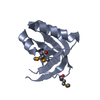
Entry Database : PDB / ID : 6xa6Title Crystal Structure of Human Scribble PDZ3:Vangl2 Complex Protein scribble homolog Vang-like protein 2 Keywords / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Homo sapiens (human)Method / / / Resolution : 1.95 Å Authors How, J.Y. / Kvansakul, M. Funding support Organization Grant number Country National Health and Medical Research Council (NHMRC, Australia) APP1103871 National Health and Medical Research Council (NHMRC, Australia) APP1079133 Australian Research Council (ARC) FT130101349
Journal : Biochem.J. / Year : 2021Title : Structural basis of the human Scribble-Vangl2 association in health and disease.Authors : How, J.Y. / Stephens, R.K. / Lim, K.Y.B. / Humbert, P.O. / Kvansakul, M. History Deposition Jun 4, 2020 Deposition site / Processing site Revision 1.0 Mar 24, 2021 Provider / Type Revision 1.1 Apr 21, 2021 Group / Category / citation_authorItem _citation.journal_volume / _citation.page_first ... _citation.journal_volume / _citation.page_first / _citation.page_last / _citation_author.name Revision 1.2 Oct 18, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession
Show all Show less
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Homo sapiens (human)
Homo sapiens (human) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.95 Å
MOLECULAR REPLACEMENT / Resolution: 1.95 Å  Authors
Authors Australia, 3items
Australia, 3items  Citation
Citation Journal: Biochem.J. / Year: 2021
Journal: Biochem.J. / Year: 2021 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 6xa6.cif.gz
6xa6.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb6xa6.ent.gz
pdb6xa6.ent.gz PDB format
PDB format 6xa6.json.gz
6xa6.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/xa/6xa6
https://data.pdbj.org/pub/pdb/validation_reports/xa/6xa6 ftp://data.pdbj.org/pub/pdb/validation_reports/xa/6xa6
ftp://data.pdbj.org/pub/pdb/validation_reports/xa/6xa6



 Links
Links Assembly
Assembly


 Components
Components Homo sapiens (human) / Gene: SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL / Production host:
Homo sapiens (human) / Gene: SCRIB, CRIB1, KIAA0147, LAP4, SCRB1, VARTUL / Production host: 
 Homo sapiens (human) / References: UniProt: Q9ULK5
Homo sapiens (human) / References: UniProt: Q9ULK5 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  Australian Synchrotron
Australian Synchrotron  / Beamline: MX1 / Wavelength: 0.9357 Å
/ Beamline: MX1 / Wavelength: 0.9357 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj











