[English] 日本語
 Yorodumi
Yorodumi- PDB-6wa1: Dimeric form of the trans-stabilized Hemolysin II C-terminal domain -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6wa1 | ||||||
|---|---|---|---|---|---|---|---|
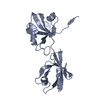
| Title | Dimeric form of the trans-stabilized Hemolysin II C-terminal domain | ||||||
 Components Components | Hemolysin II | ||||||
 Keywords Keywords | TOXIN / domain swapped dimer | ||||||
| Function / homology | Bi-component toxin, staphylococci / Leukocidin/Hemolysin toxin / Leukocidin/Hemolysin toxin family / Leukocidin/porin MspA superfamily / cytolysis in another organism / extracellular region / Hemolysin II Function and homology information Function and homology information | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR / simulated annealing | ||||||
 Authors Authors | Kaplan, A.R. / Alexandrescu, A.T. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2021 Journal: Protein Sci. / Year: 2021Title: Protein yoga: Conformational versatility of the Hemolysin II C-terminal domain detailed by NMR structures for multiple states. Authors: Kaplan, A.R. / Olson, R. / Alexandrescu, A.T. #1:  Journal: Sci Rep / Year: 2017 Journal: Sci Rep / Year: 2017Title: NMR structure of the Bacillus cereus hemolysin II C-terminal domain reveals a novel fold. Authors: Kaplan, A.R. / Kaus, K. / De, S. / Olson, R. / Alexandrescu, A.T. #2: Journal: Biomol.Nmr Assign. / Year: 2014 Title: NMR assignments for the cis and trans forms of the hemolysin II C-terminal domain. Authors: Kaplan, A.R. / Maciejewski, M.W. / Olson, R. / Alexandrescu, A.T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6wa1.cif.gz 6wa1.cif.gz | 1.4 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6wa1.ent.gz pdb6wa1.ent.gz | 1.2 MB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6wa1.json.gz 6wa1.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/wa/6wa1 https://data.pdbj.org/pub/pdb/validation_reports/wa/6wa1 ftp://data.pdbj.org/pub/pdb/validation_reports/wa/6wa1 ftp://data.pdbj.org/pub/pdb/validation_reports/wa/6wa1 | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data | |
| Other databases |
|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 10461.706 Da / Num. of mol.: 2 / Fragment: C-terminal domain (UNP residues 319-412) / Mutation: P87M Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Strain: ATCC 14579 / DSM 31 / JCM 2152 / NBRC 15305 / NCIMB 9373 / NRRL B-3711 Gene: BC_3523 / Plasmid: pET28b / Production host:  |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR | ||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR experiment |
|
- Sample preparation
Sample preparation
| Details | Type: solution Contents: 0.8 mM [U-99% 13C; U-99% 15N] HlyIIC, 20 mM sodium phosphate, 1 mM EDTA, 0.05 % w/v sodium azide, 1 mM AEBSF protease inhibitor, 90% H2O/10% D2O Label: 13C_15N_P87M / Solvent system: 90% H2O/10% D2O | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Sample |
| ||||||||||||||||||||||||
| Sample conditions | Ionic strength: 0 M / Label: 13C_15N_P87M / pH: 6.5 / Pressure: 1 atm / Temperature: 303.15 K |
-NMR measurement
| NMR spectrometer | Type: Varian INOVA / Manufacturer: Varian / Model: INOVA / Field strength: 600 MHz |
|---|
- Processing
Processing
| NMR software |
| ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method: simulated annealing / Software ordinal: 8 | ||||||||||||||||||||||||||||||||||||
| NMR representative | Selection criteria: closest to the average | ||||||||||||||||||||||||||||||||||||
| NMR ensemble | Conformer selection criteria: structures with the least restraint violations Conformers calculated total number: 100 / Conformers submitted total number: 25 |
 Movie
Movie Controller
Controller











 PDBj
PDBj
 HSQC
HSQC