[English] 日本語
 Yorodumi
Yorodumi- PDB-6uov: Cryo-EM reconstruction of the PrgHK periplasmic ring from Salmone... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6uov | ||||||
|---|---|---|---|---|---|---|---|
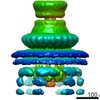
| Title | Cryo-EM reconstruction of the PrgHK periplasmic ring from Salmonella's needle complex assembled in the absence of the export apparatus | ||||||
 Components Components |
| ||||||
 Keywords Keywords | MEMBRANE PROTEIN / BACTERIAL NANOMACHINE / TYPE III SECRETION SYSTEM | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) | ||||||
| Method | ELECTRON MICROSCOPY / single particle reconstruction / cryo EM / Resolution: 3.5 Å | ||||||
 Authors Authors | Butan, C. / Galan, J. | ||||||
| Funding support |  United States, 1items United States, 1items
| ||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2019 Journal: Proc Natl Acad Sci U S A / Year: 2019Title: High-resolution view of the type III secretion export apparatus in situ reveals membrane remodeling and a secretion pathway. Authors: Carmen Butan / Maria Lara-Tejero / Wenwei Li / Jun Liu / Jorge E Galán /  Abstract: Type III protein secretion systems are essential virulence factors for many important pathogenic bacteria. The entire protein secretion machine is composed of several substructures that organize into ...Type III protein secretion systems are essential virulence factors for many important pathogenic bacteria. The entire protein secretion machine is composed of several substructures that organize into a holostructure or injectisome. The core component of the injectisome is the needle complex, which houses the export apparatus that serves as a gate for the passage of the secreted proteins through the bacterial inner membrane. Here, we describe a high-resolution structure of the export apparatus of the type III secretion system in association with the needle complex and the underlying bacterial membrane, both in isolation and in situ. We show the precise location of the core export apparatus components within the injectisome and bacterial envelope and demonstrate that their deployment results in major membrane remodeling and thinning, which may be central for the protein translocation process. We also show that InvA, a critical export apparatus component, forms a multiring cytoplasmic conduit that provides a pathway for the type III secretion substrates to reach the entrance of the export gate. Combined with structure-guided mutagenesis, our studies provide major insight into potential mechanisms of protein translocation and injectisome assembly. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6uov.cif.gz 6uov.cif.gz | 1.5 MB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6uov.ent.gz pdb6uov.ent.gz | 1.2 MB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6uov.json.gz 6uov.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/uo/6uov https://data.pdbj.org/pub/pdb/validation_reports/uo/6uov ftp://data.pdbj.org/pub/pdb/validation_reports/uo/6uov ftp://data.pdbj.org/pub/pdb/validation_reports/uo/6uov | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  20833MC  6uotC M: map data used to model this data C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
|
|---|---|
| 1 |
|
- Components
Components
| #1: Protein | Mass: 28245.287 Da / Num. of mol.: 23 / Source method: isolated from a natural source Source: (natural)  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria)References: UniProt: P41786 #2: Protein | Mass: 44509.367 Da / Num. of mol.: 23 / Source method: isolated from a natural source Source: (natural)  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria)References: UniProt: P41783 |
|---|
-Experimental details
-Experiment
| Experiment | Method: ELECTRON MICROSCOPY |
|---|---|
| EM experiment | Aggregation state: PARTICLE / 3D reconstruction method: single particle reconstruction |
- Sample preparation
Sample preparation
| Component | Name: Complex of the periplasmic domains of PrgH and PrgK from Salmonella's needle complex assembled in the absence of the export apparatus Type: COMPLEX / Entity ID: all / Source: NATURAL |
|---|---|
| Source (natural) | Organism:  Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) Salmonella enterica subsp. enterica serovar Typhimurium (bacteria) |
| Buffer solution | pH: 7.5 |
| Specimen | Embedding applied: NO / Shadowing applied: NO / Staining applied: NO / Vitrification applied: YES |
| Vitrification | Instrument: FEI VITROBOT MARK IV / Cryogen name: ETHANE |
- Electron microscopy imaging
Electron microscopy imaging
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
|---|---|
| Microscopy | Model: FEI TITAN KRIOS |
| Electron gun | Electron source:  FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM FIELD EMISSION GUN / Accelerating voltage: 300 kV / Illumination mode: FLOOD BEAM |
| Electron lens | Mode: BRIGHT FIELD |
| Image recording | Electron dose: 51 e/Å2 / Film or detector model: FEI FALCON III (4k x 4k) |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| EM software |
| ||||||||||||||||||||||||||||||||
| CTF correction | Type: PHASE FLIPPING AND AMPLITUDE CORRECTION | ||||||||||||||||||||||||||||||||
| Symmetry | Point symmetry: C23 (23 fold cyclic) | ||||||||||||||||||||||||||||||||
| 3D reconstruction | Resolution: 3.5 Å / Resolution method: FSC 0.143 CUT-OFF / Num. of particles: 10674 / Symmetry type: POINT | ||||||||||||||||||||||||||||||||
| Refinement | Cross valid method: THROUGHOUT | ||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 120.47 Å2 / Biso mean: 66.4954 Å2 / Biso min: 31.37 Å2 |
 Movie
Movie Controller
Controller









 PDBj
PDBj




