[English] 日本語
 Yorodumi
Yorodumi- PDB-6r64: N-terminal domain of modification dependent EcoKMcrA restriction ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6r64 | ||||||
|---|---|---|---|---|---|---|---|
| Title | N-terminal domain of modification dependent EcoKMcrA restriction endonuclease (NEco) in complex with C5mCGG target sequence | ||||||
 Components Components |
| ||||||
 Keywords Keywords | HYDROLASE / EcoKMcrA / NEco / N-TERMINAL DOMAIN / MODIFICATION DEPENDENT RESTRICTION / 5-METHYLCYTOSINE / 5MC / 5-HYDROXYMETHYLCYTOSINE / 5HMC / HNH ENDONUCLEASE / BBA-ME NUCLEASE / ScoMcrA | ||||||
| Function / homology |  Function and homology information Function and homology informationHydrolases; Acting on ester bonds; Endodeoxyribonucleases producing 5'-phosphomonoesters / DNA restriction-modification system / methyl-CpG binding / endonuclease activity / hydrolase activity / zinc ion binding Similarity search - Function | ||||||
| Biological species |  synthetic construct (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.64 Å MOLECULAR REPLACEMENT / Resolution: 2.64 Å | ||||||
 Authors Authors | Slyvka, A. / Zagorskaite, E. / Czapinska, H. / Sasnauskas, G. / Bochtler, M. | ||||||
 Citation Citation |  Journal: Nucleic Acids Res. / Year: 2019 Journal: Nucleic Acids Res. / Year: 2019Title: Crystal structure of the EcoKMcrA N-terminal domain (NEco): recognition of modified cytosine bases without flipping. Authors: Slyvka, A. / Zagorskaite, E. / Czapinska, H. / Sasnauskas, G. / Bochtler, M. #1:  Journal: Nucleic Acids Res. / Year: 2018 Journal: Nucleic Acids Res. / Year: 2018Title: Activity and structure of EcoKMcrA. Authors: Czapinska, H. / Kowalska, M. / Zagorskaite, E. / Manakova, E. / Slyvka, A. / Xu, S.Y. / Siksnys, V. / Sasnauskas, G. / Bochtler, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6r64.cif.gz 6r64.cif.gz | 99.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6r64.ent.gz pdb6r64.ent.gz | 72.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6r64.json.gz 6r64.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/r6/6r64 https://data.pdbj.org/pub/pdb/validation_reports/r6/6r64 ftp://data.pdbj.org/pub/pdb/validation_reports/r6/6r64 ftp://data.pdbj.org/pub/pdb/validation_reports/r6/6r64 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6t21C  6t22C  6ghcS S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 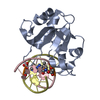
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 17368.604 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   References: UniProt: P24200, Hydrolases; Acting on ester bonds; Endodeoxyribonucleases producing 5'-phosphomonoesters #2: DNA chain | Mass: 3009.994 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) #3: DNA chain | Mass: 3108.069 Da / Num. of mol.: 2 / Source method: obtained synthetically / Source: (synth.) synthetic construct (others) #4: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 8.29 Å3/Da / Density % sol: 85.17 % |
|---|---|
| Crystal grow | Temperature: 294 K / Method: vapor diffusion, hanging drop / pH: 6.5 Details: NEco in the crystallization buffer (20 mM Tris-HCl, pH 8.0, 0.1 M NaCl, 1 mM EDTA, 2 mM DTT) was concentrated to 8.7 mg/ml and mixed in the 1:1.2 ratio with 10-mer TCAC5mCGGTTC ...Details: NEco in the crystallization buffer (20 mM Tris-HCl, pH 8.0, 0.1 M NaCl, 1 mM EDTA, 2 mM DTT) was concentrated to 8.7 mg/ml and mixed in the 1:1.2 ratio with 10-mer TCAC5mCGGTTC oligonucleotide, annealed to its complementary GAAC5mCGTGA strand. Crystals were grown by mixing 1.8 ul of the protein-DNA mixture with 2.2 ul of the condition F1 of the PACT premier crystal screen (MDL) (0.2 M NaF, 0.1 M Bis-Tris propane, pH 6.5, 20% PEG3350). Crystals were cryo-protected by the addition of 25% glycerol. |
-Data collection
| Diffraction | Mean temperature: 100 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  BESSY BESSY  / Beamline: 14.1 / Wavelength: 0.9184 Å / Beamline: 14.1 / Wavelength: 0.9184 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: Jul 10, 2018 |
| Radiation | Monochromator: KMC-1 / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9184 Å / Relative weight: 1 |
| Reflection | Resolution: 2.64→45.55 Å / Num. obs: 17671 / % possible obs: 99.8 % / Redundancy: 16 % / Biso Wilson estimate: 50.9 Å2 / CC1/2: 0.997 / Rrim(I) all: 0.23 / Rsym value: 0.223 / Net I/σ(I): 12.52 |
| Reflection shell | Resolution: 2.64→2.8 Å / Redundancy: 16.7 % / Mean I/σ(I) obs: 1.97 / Num. unique obs: 2761 / CC1/2: 0.764 / Rrim(I) all: 1.509 / Rsym value: 1.463 / % possible all: 99.2 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 6GHC Resolution: 2.64→45.55 Å / Cor.coef. Fo:Fc: 0.953 / Cor.coef. Fo:Fc free: 0.926 / SU B: 11.193 / SU ML: 0.219 / Cross valid method: THROUGHOUT / ESU R: 0.492 / ESU R Free: 0.273 / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 53.738 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.64→45.55 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller










 PDBj
PDBj
