+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 6ako | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Crystal Structure of FOXC2 DBD Bound to DBE2 DNA | |||||||||
 Components Components |
| |||||||||
 Keywords Keywords | DNA BINDING PROTEIN/DNA / FOXC / DNA binding domain / DNA recognition / Crystal Structure / Lymphoedema distichiasis syndrome / DNA BINDING PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationapoptotic process involved in outflow tract morphogenesis / negative regulation of apoptotic process involved in outflow tract morphogenesis / paraxial mesodermal cell fate commitment / positive regulation of vascular wound healing / Formation of intermediate mesoderm / glomerular endothelium development / embryonic viscerocranium morphogenesis / Formation of the ureteric bud / lymphangiogenesis / podocyte differentiation ...apoptotic process involved in outflow tract morphogenesis / negative regulation of apoptotic process involved in outflow tract morphogenesis / paraxial mesodermal cell fate commitment / positive regulation of vascular wound healing / Formation of intermediate mesoderm / glomerular endothelium development / embryonic viscerocranium morphogenesis / Formation of the ureteric bud / lymphangiogenesis / podocyte differentiation / glomerular mesangial cell development / neural crest cell development / metanephros development / regulation of organ growth / camera-type eye development / embryonic heart tube development / collagen fibril organization / positive regulation of cell adhesion mediated by integrin / artery morphogenesis / negative regulation of cold-induced thermogenesis / ureteric bud development / positive regulation of cell migration involved in sprouting angiogenesis / branching involved in blood vessel morphogenesis / ventricular cardiac muscle tissue morphogenesis / DNA-binding transcription activator activity / mesoderm development / vascular endothelial growth factor receptor signaling pathway / anatomical structure morphogenesis / blood vessel remodeling / cardiac muscle cell proliferation / somitogenesis / Notch signaling pathway / response to hormone / positive regulation of endothelial cell migration / ossification / blood vessel diameter maintenance / RNA polymerase II transcription regulatory region sequence-specific DNA binding / promoter-specific chromatin binding / chromatin DNA binding / sequence-specific double-stranded DNA binding / insulin receptor signaling pathway / heart development / DNA-binding transcription activator activity, RNA polymerase II-specific / sequence-specific DNA binding / DNA-binding transcription factor activity, RNA polymerase II-specific / cell differentiation / transcription cis-regulatory region binding / nuclear body / RNA polymerase II cis-regulatory region sequence-specific DNA binding / regulation of transcription by RNA polymerase II / positive regulation of DNA-templated transcription / chromatin / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / nucleoplasm / identical protein binding / nucleus Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.396 Å MOLECULAR REPLACEMENT / Resolution: 2.396 Å | |||||||||
 Authors Authors | Chen, X. / Wei, H. / Li, J. / Liang, X. / Dai, S. / Jiang, L. / Guo, M. / Chen, Y. | |||||||||
| Funding support |  China, 2items China, 2items
| |||||||||
 Citation Citation |  Journal: Nucleic Acids Res. / Year: 2019 Journal: Nucleic Acids Res. / Year: 2019Title: Structural basis for DNA recognition by FOXC2. Authors: Chen, X. / Wei, H. / Li, J. / Liang, X. / Dai, S. / Jiang, L. / Guo, M. / Qu, L. / Chen, Z. / Chen, L. / Chen, Y. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  6ako.cif.gz 6ako.cif.gz | 94.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb6ako.ent.gz pdb6ako.ent.gz | 67.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  6ako.json.gz 6ako.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ak/6ako https://data.pdbj.org/pub/pdb/validation_reports/ak/6ako ftp://data.pdbj.org/pub/pdb/validation_reports/ak/6ako ftp://data.pdbj.org/pub/pdb/validation_reports/ak/6ako | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  6akpC  5x07S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 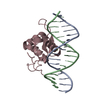
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: DNA chain | Mass: 4932.272 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  |
|---|---|
| #2: DNA chain | Mass: 4860.159 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Production host: Homo sapiens (human) / Production host:  |
| #3: Protein | Mass: 12330.169 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: FOXC2, FKHL14, MFH1 / Production host: Homo sapiens (human) / Gene: FOXC2, FKHL14, MFH1 / Production host:  |
| #4: Chemical | ChemComp-MG / |
| #5: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.47 Å3/Da / Density % sol: 64.57 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, hanging drop Details: 50 mM Bis-Tris propane (pH 6.68), 14% PEG4K, 200 mM NaCl, 10 mM MgCl2, 1 mM TCEP |
-Data collection
| Diffraction | Mean temperature: 85 K / Serial crystal experiment: N |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: Cu FINE FOCUS / Wavelength: 1.587 Å ROTATING ANODE / Type: Cu FINE FOCUS / Wavelength: 1.587 Å |
| Detector | Type: RIGAKU / Detector: CCD / Date: Mar 6, 2016 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.587 Å / Relative weight: 1 |
| Reflection | Resolution: 2.396→40.97 Å / Num. obs: 11726 / % possible obs: 92 % / Redundancy: 12.1 % / CC1/2: 1 / Rmerge(I) obs: 0.04052 / Net I/σ(I): 34.65 |
| Reflection shell | Resolution: 2.396→2.482 Å / Rmerge(I) obs: 0.3016 / Num. unique obs: 780 / CC1/2: 0.979 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 5X07 Resolution: 2.396→40.97 Å / SU ML: 0.26 / Cross valid method: FREE R-VALUE / σ(F): 1.34 / Phase error: 36.5
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.396→40.97 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj









































