+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5v5z | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
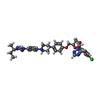
| Title | Structure of CYP51 from the pathogen Candida albicans | |||||||||
 Components Components | Lanosterol 14-alpha demethylase | |||||||||
 Keywords Keywords | OXIDOREDUCTASE/OXIDOREDUCATSE INHIBITOR / pathogen / Candida albicans / CYP51 / itraconazole / OXIDOREDUCTASE-OXIDOREDUCATSE INHIBITOR complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationcell growth mode switching, budding to filamentous / membrane raft polarization / sterol 14alpha-demethylase / ergosterol biosynthetic process / sterol 14-demethylase activity / perinuclear endoplasmic reticulum / cortical endoplasmic reticulum / endosome organization / cellular response to xenobiotic stimulus / oxidoreductase activity ...cell growth mode switching, budding to filamentous / membrane raft polarization / sterol 14alpha-demethylase / ergosterol biosynthetic process / sterol 14-demethylase activity / perinuclear endoplasmic reticulum / cortical endoplasmic reticulum / endosome organization / cellular response to xenobiotic stimulus / oxidoreductase activity / iron ion binding / heme binding / endoplasmic reticulum membrane / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |  Candida albicans (yeast) Candida albicans (yeast) | |||||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.9 Å MOLECULAR REPLACEMENT / Resolution: 2.9 Å | |||||||||
 Authors Authors | Keniya, M.V. / Sabherwal, M. / Wilson, R.K. / Sagatova, A.A. / Tyndall, J.D.A. / Monk, B.C. | |||||||||
| Funding support |  New Zealand, 2items New Zealand, 2items
| |||||||||
 Citation Citation |  Journal: Antimicrob.Agents Chemother. / Year: 2018 Journal: Antimicrob.Agents Chemother. / Year: 2018Title: Crystal Structures of Full-Length Lanosterol 14 alpha-Demethylases of Prominent Fungal Pathogens Candida albicans and Candida glabrata Provide Tools for Antifungal Discovery. Authors: Keniya, M.V. / Sabherwal, M. / Wilson, R.K. / Woods, M.A. / Sagatova, A.A. / Tyndall, J.D.A. / Monk, B.C. #1:  Journal: Antimicrob. Agents Chemother. / Year: 2015 Journal: Antimicrob. Agents Chemother. / Year: 2015Title: Structural Insights into Binding of the Antifungal Drug Fluconazole to Saccharomyces cerevisiae Lanosterol 14alpha-Demethylase. Authors: Sagatova, A.A. / Keniya, M.V. / Wilson, R.K. / Monk, B.C. / Tyndall, J.D. #2:  Journal: Proc. Natl. Acad. Sci. U.S.A. / Year: 2014 Journal: Proc. Natl. Acad. Sci. U.S.A. / Year: 2014Title: Architecture of a single membrane spanning cytochrome P450 suggests constraints that orient the catalytic domain relative to a bilayer. Authors: Monk, B.C. / Tomasiak, T.M. / Keniya, M.V. / Huschmann, F.U. / Tyndall, J.D. / O'Connell, J.D. / Cannon, R.D. / McDonald, J.G. / Rodriguez, A. / Finer-Moore, J.S. / Stroud, R.M. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5v5z.cif.gz 5v5z.cif.gz | 118.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5v5z.ent.gz pdb5v5z.ent.gz | 89.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5v5z.json.gz 5v5z.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/v5/5v5z https://data.pdbj.org/pub/pdb/validation_reports/v5/5v5z ftp://data.pdbj.org/pub/pdb/validation_reports/v5/5v5z ftp://data.pdbj.org/pub/pdb/validation_reports/v5/5v5z | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5jlcC  4lxjS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 61848.430 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Candida albicans (strain SC5314 / ATCC MYA-2876) (yeast) Candida albicans (strain SC5314 / ATCC MYA-2876) (yeast)Strain: SC5314 / ATCC MYA-2876 / Gene: ERG11, CYP51, ERG16, CAALFM_C500660CA, CaO19.922 / Production host:  |
|---|---|
| #2: Chemical | ChemComp-HEM / |
| #3: Chemical | ChemComp-1YN / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.46 Å3/Da / Density % sol: 64.42 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, hanging drop / pH: 8.8 / Details: 0.5 M GLYCINE-NAOH, 35% PEG400, |
-Data collection
| Diffraction | Mean temperature: 91 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Australian Synchrotron Australian Synchrotron  / Beamline: MX2 / Wavelength: 0.954 Å / Beamline: MX2 / Wavelength: 0.954 Å |
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Feb 21, 2015 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.954 Å / Relative weight: 1 |
| Reflection | Resolution: 2.9→91.94 Å / Num. obs: 19796 / % possible obs: 99.9 % / Redundancy: 7.5 % / CC1/2: 0.996 / Rmerge(I) obs: 0.2 / Net I/σ(I): 8.6 |
| Reflection shell | Resolution: 2.9→3.08 Å / Redundancy: 7.6 % / Rmerge(I) obs: 1.235 / Mean I/σ(I) obs: 1.7 / Num. unique obs: 3130 / CC1/2: 0.799 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4LXJ Resolution: 2.9→44.505 Å / SU ML: 0.44 / Cross valid method: FREE R-VALUE / σ(F): 1.34 / Phase error: 32.23 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.9→44.505 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller














 PDBj
PDBj