[English] 日本語
 Yorodumi
Yorodumi- PDB-5b8d: Crystal structure of a low occupancy fragment candidate (N-(4-Met... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 5b8d | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of a low occupancy fragment candidate (N-(4-Methyl-1,3-thiazol-2-yl)propanamide) bound adjacent to the ubiquitin binding pocket of the HDAC6 zinc-finger domain | ||||||
 Components Components | Histone deacetylase 6 | ||||||
 Keywords Keywords | HYDROLASE / HISTONE DEACETYLASE / HDAC / HDAC6 / FRAGMENT SCREENING / STRUCTURAL GENOMICS CONSORTIUM / SGC / DIAMOND I04-1 XCHEM / PANDDA | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of hydrogen peroxide metabolic process / cellular response to topologically incorrect protein / polyubiquitinated misfolded protein transport / negative regulation of aggrephagy / response to misfolded protein / positive regulation of protein oligomerization / type 2 mitophagy / negative regulation of protein-containing complex disassembly / peroxidase inhibitor activity / regulation of autophagy of mitochondrion ...negative regulation of hydrogen peroxide metabolic process / cellular response to topologically incorrect protein / polyubiquitinated misfolded protein transport / negative regulation of aggrephagy / response to misfolded protein / positive regulation of protein oligomerization / type 2 mitophagy / negative regulation of protein-containing complex disassembly / peroxidase inhibitor activity / regulation of autophagy of mitochondrion / erythrocyte enucleation / tubulin deacetylation / protein-containing complex disassembly / Cilium Assembly / collateral sprouting / regulation of establishment of protein localization / deacetylase activity / Transcriptional regulation by RUNX2 / tubulin deacetylase activity / lysosome localization / negative regulation of microtubule depolymerization / ATPase inhibitor activity / cilium disassembly / protein deacetylation / positive regulation of type 2 mitophagy / misfolded protein binding / dendritic spine morphogenesis / ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway / cytoplasmic ubiquitin ligase complex / regulation of androgen receptor signaling pathway / aggresome assembly / Transferases; Acyltransferases; Aminoacyltransferases / regulation of fat cell differentiation / histone deacetylase activity / cellular response to misfolded protein / protein lysine deacetylase activity / Hydrolases; Acting on carbon-nitrogen bonds, other than peptide bonds; In linear amides / aggresome / microtubule associated complex / positive regulation of intracellular estrogen receptor signaling pathway / Notch-HLH transcription pathway / negative regulation of gene expression, epigenetic / axonal transport of mitochondrion / RUNX2 regulates osteoblast differentiation / regulation of microtubule-based movement / histone deacetylase complex / cell leading edge / protein quality control for misfolded or incompletely synthesized proteins / dynein complex binding / positive regulation of epithelial cell migration / cilium assembly / regulation of macroautophagy / HSF1 activation / polyubiquitin modification-dependent protein binding / alpha-tubulin binding / negative regulation of protein-containing complex assembly / beta-tubulin binding / negative regulation of proteolysis / multivesicular body / axon cytoplasm / inclusion body / transcription corepressor binding / regulation of autophagy / actin filament organization / ubiquitin binding / regulation of protein stability / Late endosomal microautophagy / intracellular protein transport / Hsp90 protein binding / caveola / beta-catenin binding / NOTCH1 Intracellular Domain Regulates Transcription / protein destabilization / tau protein binding / Constitutive Signaling by NOTCH1 PEST Domain Mutants / Constitutive Signaling by NOTCH1 HD+PEST Domain Mutants / histone deacetylase binding / epidermal growth factor receptor signaling pathway / cellular response to hydrogen peroxide / Chaperone Mediated Autophagy / Aggrephagy / protein polyubiquitination / ubiquitin protein ligase activity / cellular response to heat / actin binding / microtubule binding / microtubule / perikaryon / ciliary basal body / RNA polymerase II cis-regulatory region sequence-specific DNA binding / axon / negative regulation of DNA-templated transcription / ubiquitin protein ligase binding / centrosome / dendrite / perinuclear region of cytoplasm / enzyme binding / zinc ion binding / nucleoplasm / nucleus Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / Resolution: 1.05 Å SYNCHROTRON / Resolution: 1.05 Å | ||||||
 Authors Authors | Harding, R.J. / Tempel, W. / Collins, P. / Pearce, N. / Brandao-Neto, J. / Douangamath, A. / Ravichandran, M. / Schapira, M. / Bountra, C. / Edwards, A.M. ...Harding, R.J. / Tempel, W. / Collins, P. / Pearce, N. / Brandao-Neto, J. / Douangamath, A. / Ravichandran, M. / Schapira, M. / Bountra, C. / Edwards, A.M. / von Delft, F. / Santhakumar, V. / Arrowsmith, C.M. / Structural Genomics Consortium (SGC) | ||||||
 Citation Citation |  Journal: J. Med. Chem. / Year: 2017 Journal: J. Med. Chem. / Year: 2017Title: Small Molecule Antagonists of the Interaction between the Histone Deacetylase 6 Zinc-Finger Domain and Ubiquitin. Authors: Harding, R.J. / Ferreira de Freitas, R. / Collins, P. / Franzoni, I. / Ravichandran, M. / Ouyang, H. / Juarez-Ornelas, K.A. / Lautens, M. / Schapira, M. / von Delft, F. / Santhakumar, V. / Arrowsmith, C.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  5b8d.cif.gz 5b8d.cif.gz | 65.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb5b8d.ent.gz pdb5b8d.ent.gz | 46.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  5b8d.json.gz 5b8d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/b8/5b8d https://data.pdbj.org/pub/pdb/validation_reports/b8/5b8d ftp://data.pdbj.org/pub/pdb/validation_reports/b8/5b8d ftp://data.pdbj.org/pub/pdb/validation_reports/b8/5b8d | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  5kh3C  5kh7C  5kh9C  5wpbC  3c5kS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 1 molecules A
| #1: Protein | Mass: 11932.607 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: HDAC6, KIAA0901, JM21 / Plasmid: pET28-lic / Production host: Homo sapiens (human) / Gene: HDAC6, KIAA0901, JM21 / Plasmid: pET28-lic / Production host:  |
|---|
-Non-polymers , 6 types, 122 molecules 

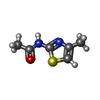








| #2: Chemical | | #3: Chemical | #4: Chemical | ChemComp-6T4 / ~{ | #5: Chemical | ChemComp-UNX / #6: Chemical | ChemComp-NA / | #7: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.17 Å3/Da / Density % sol: 43.26 % |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop / pH: 4.6 Details: 2M Na-formate, 0.2M Na-acetate pH4.6, 5% ethylene glycol |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Diamond Diamond  / Beamline: I04-1 / Wavelength: 0.92819 Å / Beamline: I04-1 / Wavelength: 0.92819 Å | ||||||||||||||||||||||||||||||
| Detector | Type: DECTRIS PILATUS 6M-F / Detector: PIXEL / Date: Sep 30, 2015 | ||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.92819 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.05→34.8 Å / Num. obs: 47523 / % possible obs: 97.4 % / Redundancy: 5.4 % / CC1/2: 0.999 / Rmerge(I) obs: 0.058 / Rpim(I) all: 0.026 / Rrim(I) all: 0.064 / Net I/σ(I): 14.6 / Num. measured all: 257744 | ||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1 / Rejects: _
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Starting model: pdbid 3C5K Resolution: 1.05→34.8 Å / Cor.coef. Fo:Fc: 0.98 / Cor.coef. Fo:Fc free: 0.977 / SU B: 0.733 / SU ML: 0.016 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.023 / ESU R Free: 0.023 / Stereochemistry target values: MAXIMUM LIKELIHOOD Details: Users of this crystal structure: verify our intepretion of the electron density. Amplitudes and unmerged intensities are included with this deposition. Diffraction images will be deposited ...Details: Users of this crystal structure: verify our intepretion of the electron density. Amplitudes and unmerged intensities are included with this deposition. Diffraction images will be deposited in a public repository. Geometry restraints for the fragment candidate were prepared with GRADE. The methyl group of the fragment candidate was not resolved by electron density and was omitted from the model. Ambiguous difference density suggests some form of covalent modification of Cys1147.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 69.28 Å2 / Biso mean: 13.189 Å2 / Biso min: 6.88 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: final / Resolution: 1.05→34.8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 1.052→1.08 Å / Total num. of bins used: 20
|
 Movie
Movie Controller
Controller










 PDBj
PDBj









