Entry Database : PDB / ID : 4zy3Title Crystal Structure of Keap1 in Complex with a small chemical compound, K67 Kelch-like ECH-associated protein 1 Keywords / / / / / / / Function / homology Function Domain/homology Component
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / Biological species Mus musculus (house mouse)Method / / / Resolution : 1.8 Å Authors Fukutomi, T. / Iso, T. / Suzuki, T. / Takagi, K. / Mizushima, T. / Komatsu, M. / Yamamoto, M. Journal : Nat Commun / Year : 2016Title : p62/Sqstm1 promotes malignancy of HCV-positive hepatocellular carcinoma through Nrf2-dependent metabolic reprogrammingAuthors: Saito, T. / Ichimura, Y. / Taguchi, K. / Suzuki, T. / Mizushima, T. / Takagi, K. / Hirose, Y. / Nagahashi, M. / Iso, T. / Fukutomi, T. / Ohishi, M. / Endo, K. / Uemura, T. / Nishito, Y. / ... Authors : Saito, T. / Ichimura, Y. / Taguchi, K. / Suzuki, T. / Mizushima, T. / Takagi, K. / Hirose, Y. / Nagahashi, M. / Iso, T. / Fukutomi, T. / Ohishi, M. / Endo, K. / Uemura, T. / Nishito, Y. / Okuda, S. / Obata, M. / Kouno, T. / Imamura, R. / Tada, Y. / Obata, R. / Yasuda, D. / Takahashi, K. / Fujimura, T. / Pi, J. / Lee, M.S. / Ueno, T. / Ohe, T. / Mashino, T. / Wakai, T. / Kojima, H. / Okabe, T. / Nagano, T. / Motohashi, H. / Waguri, S. / Soga, T. / Yamamoto, M. / Tanaka, K. / Komatsu, M. History Deposition May 21, 2015 Deposition site / Processing site Revision 1.0 May 25, 2016 Provider / Type Revision 1.1 Jul 13, 2016 Group Revision 1.2 Feb 19, 2020 Group / Derived calculations / Category / pdbx_struct_oper_listItem / _pdbx_struct_oper_list.symmetry_operationRevision 1.3 Nov 8, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accessionRevision 1.4 Oct 16, 2024 Group / Category / pdbx_modification_feature
Show all Show less
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.8 Å
MOLECULAR REPLACEMENT / Resolution: 1.8 Å  Authors
Authors Citation
Citation Journal: Nat Commun / Year: 2016
Journal: Nat Commun / Year: 2016 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4zy3.cif.gz
4zy3.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4zy3.ent.gz
pdb4zy3.ent.gz PDB format
PDB format 4zy3.json.gz
4zy3.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/zy/4zy3
https://data.pdbj.org/pub/pdb/validation_reports/zy/4zy3 ftp://data.pdbj.org/pub/pdb/validation_reports/zy/4zy3
ftp://data.pdbj.org/pub/pdb/validation_reports/zy/4zy3
 Links
Links Assembly
Assembly

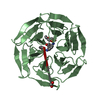
 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  SPring-8
SPring-8  / Beamline: BL44XU / Wavelength: 0.9 Å
/ Beamline: BL44XU / Wavelength: 0.9 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller












 PDBj
PDBj








