| Entry | Database: PDB / ID: 4qpw
|
|---|
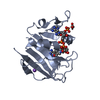
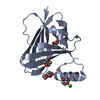
| Title | BiXyn10A CBM1 with Xylohexaose Bound |
|---|
 Components Components | glycosyl hydrolase family 10 |
|---|
 Keywords Keywords | HYDROLASE / Carbohydrate-Binding Module (CBM) |
|---|
| Function / homology |  Function and homology information Function and homology information
Glycoside hydrolase family 10 / Glycoside hydrolase family 10 domain / Glycosyl hydrolase family 10 / Glycosyl hydrolase family 10 / Galactose-binding domain-like / Galactose-binding-like domain superfamily / Prokaryotic membrane lipoprotein lipid attachment site profile. / Glycoside hydrolase superfamily / Jelly Rolls / Sandwich / Mainly BetaSimilarity search - Domain/homology |
|---|
| Biological species |  Bacteroides intestinalis DSM 17393 (bacteria) Bacteroides intestinalis DSM 17393 (bacteria) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.14 Å MOLECULAR REPLACEMENT / Resolution: 1.14 Å |
|---|
 Authors Authors | Chekan, J.R. / Nair, S.K. |
|---|
 Citation Citation |  Journal: Proc.Natl.Acad.Sci.USA / Year: 2014 Journal: Proc.Natl.Acad.Sci.USA / Year: 2014
Title: Xylan utilization in human gut commensal bacteria is orchestrated by unique modular organization of polysaccharide-degrading enzymes.
Authors: Zhang, M. / Chekan, J.R. / Dodd, D. / Hong, P.Y. / Radlinski, L. / Revindran, V. / Nair, S.K. / Mackie, R.I. / Cann, I. |
|---|
| History | | Deposition | Jun 25, 2014 | Deposition site: RCSB / Processing site: RCSB |
|---|
| Revision 1.0 | Aug 13, 2014 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Sep 24, 2014 | Group: Other |
|---|
| Revision 1.2 | Oct 1, 2014 | Group: Database references |
|---|
| Revision 2.0 | Jul 29, 2020 | Group: Atomic model / Data collection ...Atomic model / Data collection / Derived calculations / Structure summary
Category: atom_site / chem_comp ...atom_site / chem_comp / entity / entity_name_com / pdbx_branch_scheme / pdbx_chem_comp_identifier / pdbx_entity_branch / pdbx_entity_branch_descriptor / pdbx_entity_branch_link / pdbx_entity_branch_list / pdbx_entity_nonpoly / pdbx_molecule_features / pdbx_nonpoly_scheme / pdbx_struct_assembly_gen / struct_asym / struct_conn / struct_site / struct_site_gen
Item: _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ..._atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_asym_id / _atom_site.auth_atom_id / _atom_site.auth_seq_id / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.type_symbol / _chem_comp.name / _chem_comp.type / _entity.formula_weight / _entity.pdbx_description / _entity.pdbx_number_of_molecules / _entity.type / _pdbx_struct_assembly_gen.asym_id_list / _struct_conn.pdbx_dist_value / _struct_conn.pdbx_leaving_atom_flag / _struct_conn.ptnr1_auth_asym_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr2_auth_asym_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id
Description: Carbohydrate remediation / Provider: repository / Type: Remediation |
|---|
| Revision 2.1 | Feb 28, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Refinement description / Structure summary
Category: chem_comp / chem_comp_atom ...chem_comp / chem_comp_atom / chem_comp_bond / database_2 / software / struct_conn
Item: _chem_comp.pdbx_synonyms / _database_2.pdbx_DOI ..._chem_comp.pdbx_synonyms / _database_2.pdbx_DOI / _database_2.pdbx_database_accession / _software.name / _struct_conn.pdbx_leaving_atom_flag |
|---|
|
|---|
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information Bacteroides intestinalis DSM 17393 (bacteria)
Bacteroides intestinalis DSM 17393 (bacteria) X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.14 Å
MOLECULAR REPLACEMENT / Resolution: 1.14 Å  Authors
Authors Citation
Citation Journal: Proc.Natl.Acad.Sci.USA / Year: 2014
Journal: Proc.Natl.Acad.Sci.USA / Year: 2014 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 4qpw.cif.gz
4qpw.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb4qpw.ent.gz
pdb4qpw.ent.gz PDB format
PDB format 4qpw.json.gz
4qpw.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/qp/4qpw
https://data.pdbj.org/pub/pdb/validation_reports/qp/4qpw ftp://data.pdbj.org/pub/pdb/validation_reports/qp/4qpw
ftp://data.pdbj.org/pub/pdb/validation_reports/qp/4qpw Links
Links Assembly
Assembly
 Components
Components Bacteroides intestinalis DSM 17393 (bacteria)
Bacteroides intestinalis DSM 17393 (bacteria)
 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  APS
APS  / Beamline: 21-ID-G / Wavelength: 0.97857 Å
/ Beamline: 21-ID-G / Wavelength: 0.97857 Å Processing
Processing MOLECULAR REPLACEMENT / Resolution: 1.14→47.44 Å / Cor.coef. Fo:Fc: 0.976 / Cor.coef. Fo:Fc free: 0.971 / SU B: 0.487 / SU ML: 0.023 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.03 / ESU R Free: 0.032 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
MOLECULAR REPLACEMENT / Resolution: 1.14→47.44 Å / Cor.coef. Fo:Fc: 0.976 / Cor.coef. Fo:Fc free: 0.971 / SU B: 0.487 / SU ML: 0.023 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.03 / ESU R Free: 0.032 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS Movie
Movie Controller
Controller













 PDBj
PDBj






