[English] 日本語
 Yorodumi
Yorodumi- PDB-7dfr: CRYSTAL STRUCTURES OF ESCHERICHIA COLI DIHYDROFOLATE REDUCTASE. T... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 7dfr | ||||||
|---|---|---|---|---|---|---|---|
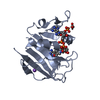
| Title | CRYSTAL STRUCTURES OF ESCHERICHIA COLI DIHYDROFOLATE REDUCTASE. THE NADP+ HOLOENZYME AND THE FOLATE(DOT)NADP+ TERNARY COMPLEX. SUBSTRATE BINDING AND A MODEL FOR THE TRANSITION STATE | ||||||
 Components Components | DIHYDROFOLATE REDUCTASE | ||||||
 Keywords Keywords | OXIDOREDUCTASE / OXIDO-REDUCTASE | ||||||
| Function / homology |  Function and homology information Function and homology informationmethotrexate binding / dihydrofolic acid binding / 10-formyltetrahydrofolate biosynthetic process / response to methotrexate / folic acid biosynthetic process / folic acid binding / NADP+ binding / dihydrofolate metabolic process / dihydrofolate reductase / dihydrofolate reductase activity ...methotrexate binding / dihydrofolic acid binding / 10-formyltetrahydrofolate biosynthetic process / response to methotrexate / folic acid biosynthetic process / folic acid binding / NADP+ binding / dihydrofolate metabolic process / dihydrofolate reductase / dihydrofolate reductase activity / folic acid metabolic process / tetrahydrofolate biosynthetic process / NADPH binding / one-carbon metabolic process / NADP binding / response to xenobiotic stimulus / response to antibiotic / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.5 Å X-RAY DIFFRACTION / Resolution: 2.5 Å | ||||||
 Authors Authors | Bystroff, C. / Oatley, S.J. / Kraut, J. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1990 Journal: Biochemistry / Year: 1990Title: Crystal structures of Escherichia coli dihydrofolate reductase: the NADP+ holoenzyme and the folate.NADP+ ternary complex. Substrate binding and a model for the transition state. Authors: Bystroff, C. / Oatley, S.J. / Kraut, J. #1:  Journal: To be Published Journal: To be PublishedTitle: Crystal Structure of Unliganded Escherichia Coli Dihydrofolate Reductase. Ligand-Induced Conformational Changes and Cooperativity in Binding Authors: Bystroff, C. / Kraut, J. #2:  Journal: J.Biol.Chem. / Year: 1982 Journal: J.Biol.Chem. / Year: 1982Title: Crystal Structures of Escherichia Coli and Lactobacillus Casei Dihydrofolate Reductase Refined at 1.7 Angstroms Resolution. I. General Features and Binding of Methotrexate Authors: Bolin, J.T. / Filman, D.J. / Matthews, D.A. / Hamlin, R.C. / Kraut, J. #3:  Journal: Biochemistry / Year: 1987 Journal: Biochemistry / Year: 1987Title: Effect of Single Amino Acid Replacements on the Folding and Stability of Dihydrofolate Reductase from Escherichia Coli Authors: Perry, K.M. / Onuffer, J.J. / Touchette, N.A. / Herndon, C.S. / Gittelman, M.S. / Matthews, C.R. / Chen, J.-T. / Mayer, R.J. / Taira, K. / Benkovic, S.J. / Howell, E.E. / Kraut, J. #4:  Journal: J.Biol.Chem. / Year: 1982 Journal: J.Biol.Chem. / Year: 1982Title: Crystal Structures of Escherichia Coli and Lactobacillus Casei Dihydrofolate Reductase Refined at 1.7 Angstroms Resolution. II. Environment of Bound Nadph and Implications for Catalysis Authors: Filman, D.J. / Bolin, J.T. / Matthews, D.A. / Kraut, J. #5:  Journal: J.Biol.Chem. / Year: 1982 Journal: J.Biol.Chem. / Year: 1982Title: Crystal Structure of Avian Dihydrofolate Reductase Containing Phenyltriazine and Nadph Authors: Volz, K.W. / Matthews, D.A. / Alden, R.A. / Freer, S.T. / Hansch, C. / Kaufman, B.T. / Kraut, J. #6:  Journal: Biochemistry / Year: 1979 Journal: Biochemistry / Year: 1979Title: Interpretation of Nuclear Magnetic Resonance Spectra for Lactobacillus Casei Dihydrofolate Reductase Based on the X-Ray Structure of the Enzyme-Methotrexate-Nadph Complex Authors: Matthews, D.A. #7:  Journal: J.Biol.Chem. / Year: 1979 Journal: J.Biol.Chem. / Year: 1979Title: Dihydrofolate Reductase from Lactobacillus Casei. Stereochemistry of Nadph Binding Authors: Matthews, D.A. / Alden, R.A. / Freer, S.T. / Xuong, N.-H. / Kraut, J. #8:  Journal: J.Biol.Chem. / Year: 1979 Journal: J.Biol.Chem. / Year: 1979Title: Proton Magnetic Resonance Studies on Escherichia Coli Dihydrofolate Reductase. Assignment of Histidine C-2 Protons in Binary Complexes with Folates on the Basis of the Crystal Structure with ...Title: Proton Magnetic Resonance Studies on Escherichia Coli Dihydrofolate Reductase. Assignment of Histidine C-2 Protons in Binary Complexes with Folates on the Basis of the Crystal Structure with Methotrexate and on Chemical Modifications Authors: Poe, M. / Hoogsteen, K. / Matthews, D.A. #9:  Journal: J.Biol.Chem. / Year: 1978 Journal: J.Biol.Chem. / Year: 1978Title: Dihydrofolate Reductase from Lactobacillus Casei. X-Ray Structure of the Enzyme-Methotrexate-Nadph Complex Authors: Matthews, D.A. / Alden, R.A. / Bolin, J.T. / Filman, D.J. / Freer, S.T. / Hamlin, R. / Hol, W.G.J. / Kisliuk, R.L. / Pastore, E.J. / Plante, L.T. / Xuong, N.-H. / Kraut, J. #10:  Journal: Biochemistry / Year: 1978 Journal: Biochemistry / Year: 1978Title: Dihydrofolate Reductase. The Amino Acid Sequence of the Enzyme from a Methotrexate-Resistant Mutant of Escherichia Coli Authors: Bennett, C.D. / Rodkey, J.A. / Sondey, J.M. / Hirschmann, R. #11:  Journal: Science / Year: 1977 Journal: Science / Year: 1977Title: Dihydrofolate Reductase. X-Ray Structure of the Binary Complex with Methotrexate Authors: Matthews, D.A. / Alden, R.A. / Bolin, J.T. / Freer, S.T. / Hamlin, R. / Xuong, N. / Kraut, J. / Poe, M. / Williams, M. / Hoogsteen, K. #12:  Journal: Biochemistry / Year: 1972 Journal: Biochemistry / Year: 1972Title: Dihydrofolate Reductase. Purification and Characterization of the Enzyme from an Amethopterin-Resistant Mutant of Escherichia Coli Authors: Poe, M. / Greenfield, N.J. / Hirshfield, J.M. / Williams, M.N. / Hoogsteen, K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  7dfr.cif.gz 7dfr.cif.gz | 49.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb7dfr.ent.gz pdb7dfr.ent.gz | 34.9 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  7dfr.json.gz 7dfr.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/df/7dfr https://data.pdbj.org/pub/pdb/validation_reports/df/7dfr ftp://data.pdbj.org/pub/pdb/validation_reports/df/7dfr ftp://data.pdbj.org/pub/pdb/validation_reports/df/7dfr | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||||||
| Unit cell |
| ||||||||||||
| Atom site foot note | 1: TEN HYDROPHILIC SIDE CHAINS ARE EITHER ENTIRELY OR PARTIALLY MISSING AND NO COORDINATES ARE INCLUDED FOR THEM IN THIS ENTRY - GLU 17, ARG 44, GLU 48, ARG 52, ARG 98, LYS 106, ASP 116, GLU 120, GLU 129, AND ASP 131. 2: THE PEPTIDE BOND LINKING GLY 95 TO GLY 96 IS IN THE CIS CONFORMATION. | ||||||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 18020.326 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  |
|---|---|
| #2: Chemical | ChemComp-FOL / |
| #3: Chemical | ChemComp-NAP / |
| #4: Water | ChemComp-HOH / |
| Compound details | RESIDUES 16 - 19 FORM A TYPE I BETA TURN. THIS PART OF THE STRUCTURE IS DENOTED AS THE MET 20 LOOP. |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.27 Å3/Da / Density % sol: 62.37 % | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS pH: 7 / Method: vapor diffusion, hanging drop | |||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Reflection | *PLUS Highest resolution: 2.5 Å / Num. all: 8698 / Num. obs: 8586 / Num. measured all: 68035 / Rmerge F obs: 0.049 |
|---|---|
| Reflection shell | *PLUS Highest resolution: 2.5 Å / Lowest resolution: 2.6 Å |
- Processing
Processing
| Software | Name: PROLSQ / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor obs: 0.245 / Highest resolution: 2.5 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 2.5 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.5 Å / Lowest resolution: 20 Å / Rfactor obs: 0.245 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS |
 Movie
Movie Controller
Controller












 PDBj
PDBj