[English] 日本語
 Yorodumi
Yorodumi- PDB-4mwe: Trypanosoma brucei methionyl-tRNA synthetase in complex with inhi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4mwe | ||||||
|---|---|---|---|---|---|---|---|
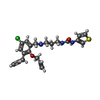
| Title | Trypanosoma brucei methionyl-tRNA synthetase in complex with inhibitor 1-(3-{[5-chloro-3-(prop-2-en-1-yl)-2-(prop-2-en-1-yloxy)benzyl]amino}propyl)-3-thiophen-3-ylurea (Chem 1475) | ||||||
 Components Components | Methionyl-tRNA synthetase | ||||||
 Keywords Keywords | LIGASE/LIGASE INHIBITOR / aminoacyl-tRNA synthetase / aaRS / MetRS / parasite / protein-inhibitor complex / Rossmann fold / translation / nucleotide binding / LIGASE-LIGASE INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology informationmethionine-tRNA ligase / methionine-tRNA ligase activity / methionyl-tRNA aminoacylation / ciliary plasm / mitochondrion / ATP binding / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / RIGID BODY REFINEMENT OF ISOMORPHOUS STRUCTURE / Resolution: 2.45 Å SYNCHROTRON / RIGID BODY REFINEMENT OF ISOMORPHOUS STRUCTURE / Resolution: 2.45 Å | ||||||
 Authors Authors | Koh, C.Y. / Kim, J.E. / Wetzel, A.B. / de van der Schueren, W.J. / Shibata, S. / Liu, J. / Zhang, Z. / Fan, E. / Verlinde, C.L.M.J. / Hol, W.G.J. | ||||||
 Citation Citation |  Journal: Plos Negl Trop Dis / Year: 2014 Journal: Plos Negl Trop Dis / Year: 2014Title: Structures of Trypanosoma brucei Methionyl-tRNA Synthetase with Urea-Based Inhibitors Provide Guidance for Drug Design against Sleeping Sickness. Authors: Koh, C.Y. / Kim, J.E. / Wetzel, A.B. / de van der Schueren, W.J. / Shibata, S. / Ranade, R.M. / Liu, J. / Zhang, Z. / Gillespie, J.R. / Buckner, F.S. / Verlinde, C.L. / Fan, E. / Hol, W.G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4mwe.cif.gz 4mwe.cif.gz | 434.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4mwe.ent.gz pdb4mwe.ent.gz | 356 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4mwe.json.gz 4mwe.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/mw/4mwe https://data.pdbj.org/pub/pdb/validation_reports/mw/4mwe ftp://data.pdbj.org/pub/pdb/validation_reports/mw/4mwe ftp://data.pdbj.org/pub/pdb/validation_reports/mw/4mwe | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4mvwC  4mvxC  4mvyC  4mw0C  4mw1C  4mw2C  4mw4C  4mw5C  4mw6C  4mw7C  4mw9C  4mwbC  4mwcC  4mwdC  4eg8S C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 2 molecules AB
| #1: Protein | Mass: 61434.707 Da / Num. of mol.: 2 / Fragment: UNP residues 237-773 / Mutation: K456A, K457R, E458A Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|
-Non-polymers , 6 types, 274 molecules 









| #2: Chemical | ChemComp-GOL / #3: Chemical | ChemComp-DMS / #4: Chemical | #5: Chemical | ChemComp-MET / | #6: Chemical | ChemComp-2EN / | #7: Water | ChemComp-HOH / | |
|---|
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.9 Å3/Da / Density % sol: 68.44 % |
|---|---|
| Crystal grow | Temperature: 298 K / Method: vapor diffusion, sitting drop Details: 2.0-2.3 M ammonium sulfate, 0.2 M sodium chloride, 0.1 M sodium cacodylate, pH 6.0-6.8, VAPOR DIFFUSION, SITTING DROP, temperature 298K PH range: 6.0-6.8 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  SSRL SSRL  / Beamline: BL12-2 / Wavelength: 0.98 Å / Beamline: BL12-2 / Wavelength: 0.98 Å |
| Detector | Type: DECTRIS PILATUS 6M / Detector: PIXEL / Date: May 20, 2011 |
| Diffraction measurement | Details: 1.00 degrees, 10.0 sec, detector distance 420.00 mm |
| Radiation | Monochromator: Liquid nitrogen-cooled double crystal Si(111) Protocol: SINGLE WAVELENGTH / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.98 Å / Relative weight: 1 |
| Reflection | Av R equivalents: 0.146 / Number: 381757 |
| Reflection | Resolution: 2.45→50 Å / Num. obs: 70116 / % possible obs: 98.8 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -3 / Redundancy: 5.4 % / Rmerge(I) obs: 0.141 / Rsym value: 0.141 / Χ2: 1.063 / Net I/σ(I): 6.6 |
| Reflection shell | Resolution: 2.45→2.49 Å / Redundancy: 5.2 % / Rmerge(I) obs: 0 / Mean I/σ(I) obs: 1.97 / Rsym value: 0 / % possible all: 99.8 |
| Cell measurement | Reflection used: 381757 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: RIGID BODY REFINEMENT OF ISOMORPHOUS STRUCTURE Starting model: PDB ENTRY 4EG8 Resolution: 2.45→30 Å / Cor.coef. Fo:Fc: 0.944 / Cor.coef. Fo:Fc free: 0.93 / WRfactor Rfree: 0.231 / WRfactor Rwork: 0.204 / Occupancy max: 1 / Occupancy min: 0.5 / SU B: 13.684 / SU ML: 0.152 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R: 0.273 / ESU R Free: 0.212 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: BABINET MODEL WITH MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 160.74 Å2 / Biso mean: 49.0462 Å2 / Biso min: 20.23 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.45→30 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 20
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller

























 PDBj
PDBj





