[English] 日本語
 Yorodumi
Yorodumi- PDB-4gr8: Crystal structure of the catalytic domain of Human MMP12 in compl... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 4gr8 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of the catalytic domain of Human MMP12 in complex with selective phosphinic inhibitor RXP470C | ||||||
 Components Components | Macrophage metalloelastase | ||||||
 Keywords Keywords | HYDROLASE/HYDROLASE INHIBITOR / potent selective phosphinic inhibitor / METZINCIN / Zinc protease / HYDROLASE-HYDROLASE INHIBITOR complex | ||||||
| Function / homology |  Function and homology information Function and homology informationmacrophage elastase / negative regulation of endothelial cell-matrix adhesion / bronchiole development / positive regulation of epithelial cell proliferation involved in wound healing / elastin catabolic process / regulation of defense response to virus by host / positive regulation of type I interferon-mediated signaling pathway / wound healing, spreading of epidermal cells / negative regulation of type I interferon-mediated signaling pathway / lung alveolus development ...macrophage elastase / negative regulation of endothelial cell-matrix adhesion / bronchiole development / positive regulation of epithelial cell proliferation involved in wound healing / elastin catabolic process / regulation of defense response to virus by host / positive regulation of type I interferon-mediated signaling pathway / wound healing, spreading of epidermal cells / negative regulation of type I interferon-mediated signaling pathway / lung alveolus development / response to amyloid-beta / Collagen degradation / collagen catabolic process / positive regulation of interferon-alpha production / extracellular matrix disassembly / core promoter sequence-specific DNA binding / collagen binding / Degradation of the extracellular matrix / extracellular matrix organization / metalloendopeptidase activity / cellular response to virus / protein import into nucleus / extracellular matrix / endopeptidase activity / sequence-specific DNA binding / serine-type endopeptidase activity / calcium ion binding / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / proteolysis / : / extracellular region / zinc ion binding / nucleus / cytoplasm Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.299 Å MOLECULAR REPLACEMENT / Resolution: 1.299 Å | ||||||
 Authors Authors | Stura, E.A. / Vera, L. / Beau, F. / Devel, L. / Cassar-Lajeunesse, E. / Dive, V. | ||||||
 Citation Citation |  Journal: J.Med.Chem. / Year: 2013 Journal: J.Med.Chem. / Year: 2013Title: Molecular determinants of a selective matrix metalloprotease-12 inhibitor: insights from crystallography and thermodynamic studies. Authors: Czarny, B. / Stura, E.A. / Devel, L. / Vera, L. / Cassar-Lajeunesse, E. / Beau, F. / Calderone, V. / Fragai, M. / Luchinat, C. / Dive, V. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  4gr8.cif.gz 4gr8.cif.gz | 54.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb4gr8.ent.gz pdb4gr8.ent.gz | 36.8 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  4gr8.json.gz 4gr8.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/gr/4gr8 https://data.pdbj.org/pub/pdb/validation_reports/gr/4gr8 ftp://data.pdbj.org/pub/pdb/validation_reports/gr/4gr8 ftp://data.pdbj.org/pub/pdb/validation_reports/gr/4gr8 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  4gqlSC  4gr0C  4gr3C C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
-Protein , 1 types, 1 molecules A
| #1: Protein | Mass: 16830.721 Da / Num. of mol.: 1 / Fragment: catalytic domain (UNP residues 111-262) / Mutation: F171D Source method: isolated from a genetically manipulated source Details: N-TERMINAL MISSING RESIDUES (106-110) ARE DUE TO SELF-PROTEOLYSIS. C-terminal glycine 263 has also been proteolyzed. Source: (gene. exp.)  Homo sapiens (human) / Gene: HME, MMP12 / Plasmid: PET24A / Production host: Homo sapiens (human) / Gene: HME, MMP12 / Plasmid: PET24A / Production host:  |
|---|
-Non-polymers , 5 types, 204 molecules 

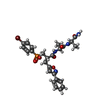






| #2: Chemical | | #3: Chemical | #4: Chemical | ChemComp-R4C / | #5: Chemical | ChemComp-IMD / | #6: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.07 Å3/Da / Density % sol: 40.64 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, sitting drop / pH: 8.5 Details: protein: hMMP12 F171D 848 microM. reservoir: 24% PEG 10000, 0.2 M imidazole malate pH 8.5. CRYOPROTECTANT: 27% PEG 8000, 15% monomethyl-PEG 550, 10% glycerol, 0.09 M Tris-HCl, pH 8.0, VAPOR ...Details: protein: hMMP12 F171D 848 microM. reservoir: 24% PEG 10000, 0.2 M imidazole malate pH 8.5. CRYOPROTECTANT: 27% PEG 8000, 15% monomethyl-PEG 550, 10% glycerol, 0.09 M Tris-HCl, pH 8.0, VAPOR DIFFUSION, SITTING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID14-4 / Wavelength: 0.9765 Å / Beamline: ID14-4 / Wavelength: 0.9765 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 315r / Detector: CCD / Date: Apr 24, 2010 / Details: mirrors | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Monochromator: Si 111 CHANNEL / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 0.9765 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 1.3→50 Å / Num. all: 35305 / Num. obs: 35086 / % possible obs: 99.4 % / Observed criterion σ(F): 0 / Observed criterion σ(I): -4 / Redundancy: 7.56 % / Biso Wilson estimate: 17.669 Å2 / Rmerge(I) obs: 0.098 / Rsym value: 0.091 / Net I/σ(I): 12.84 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell | Diffraction-ID: 1
|
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 4GQL Resolution: 1.299→30.008 Å / SU ML: 0.09 / Isotropic thermal model: Isotropic / Cross valid method: THROUGHOUT / σ(F): 1.99 / σ(I): -3 / Phase error: 19.25 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.299→30.008 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION
|
 Movie
Movie Controller
Controller
















 PDBj
PDBj













