[English] 日本語
 Yorodumi
Yorodumi- PDB-3zrz: Crystal structure of the second and third fibronectin F1 modules ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3zrz | ||||||
|---|---|---|---|---|---|---|---|
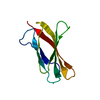
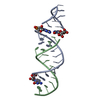
| Title | Crystal structure of the second and third fibronectin F1 modules in complex with a fragment of Streptococcus pyogenes SfbI-5 | ||||||
 Components Components |
| ||||||
 Keywords Keywords | CELL ADHESION / PRTF / BETA ZIPPER | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of monocyte activation / negative regulation of transforming growth factor beta production / Extracellular matrix organization / positive regulation of substrate-dependent cell migration, cell attachment to substrate / calcium-independent cell-matrix adhesion / neural crest cell migration involved in autonomic nervous system development / Fibronectin matrix formation / fibrinogen complex / peptide cross-linking / integrin activation ...negative regulation of monocyte activation / negative regulation of transforming growth factor beta production / Extracellular matrix organization / positive regulation of substrate-dependent cell migration, cell attachment to substrate / calcium-independent cell-matrix adhesion / neural crest cell migration involved in autonomic nervous system development / Fibronectin matrix formation / fibrinogen complex / peptide cross-linking / integrin activation / ALK mutants bind TKIs / cell-substrate junction assembly / proteoglycan binding / extracellular matrix structural constituent / Molecules associated with elastic fibres / MET activates PTK2 signaling / peptidase activator activity / Syndecan interactions / biological process involved in interaction with symbiont / p130Cas linkage to MAPK signaling for integrins / response to muscle activity / endoplasmic reticulum-Golgi intermediate compartment / endodermal cell differentiation / regulation of protein phosphorylation / GRB2:SOS provides linkage to MAPK signaling for Integrins / basement membrane / Non-integrin membrane-ECM interactions / ECM proteoglycans / Integrin cell surface interactions / endothelial cell migration / regulation of ERK1 and ERK2 cascade / positive regulation of axon extension / collagen binding / Degradation of the extracellular matrix / Integrin signaling / substrate adhesion-dependent cell spreading / platelet alpha granule lumen / Turbulent (oscillatory, disturbed) flow shear stress activates signaling by PIEZO1 and integrins in endothelial cells / cell-matrix adhesion / acute-phase response / Cell surface interactions at the vascular wall / integrin-mediated signaling pathway / Post-translational protein phosphorylation / wound healing / Signaling by high-kinase activity BRAF mutants / MAP2K and MAPK activation / : / response to wounding / integrin binding / extracellular matrix / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / positive regulation of fibroblast proliferation / Signaling by RAF1 mutants / Signaling by moderate kinase activity BRAF mutants / Paradoxical activation of RAF signaling by kinase inactive BRAF / Signaling downstream of RAS mutants / Signaling by BRAF and RAF1 fusions / Signaling by ALK fusions and activated point mutants / Platelet degranulation / nervous system development / regulation of cell shape / GPER1 signaling / heparin binding / heart development / protease binding / angiogenesis / Interleukin-4 and Interleukin-13 signaling / blood microparticle / positive regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / cell adhesion / apical plasma membrane / endoplasmic reticulum lumen / receptor ligand activity / signaling receptor binding / positive regulation of cell population proliferation / positive regulation of gene expression / extracellular space / extracellular exosome / extracellular region / identical protein binding / membrane / plasma membrane Similarity search - Function | ||||||
| Biological species |  HOMO SAPIENS (human) HOMO SAPIENS (human) STREPTOCOCCUS PYOGENES (bacteria) STREPTOCOCCUS PYOGENES (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 1.7 Å MOLECULAR REPLACEMENT / Resolution: 1.7 Å | ||||||
 Authors Authors | Norris, N.C. / Bingham, R.J. / Potts, J.R. | ||||||
 Citation Citation |  Journal: J.Biol.Chem. / Year: 2011 Journal: J.Biol.Chem. / Year: 2011Title: Structural and Functional Analysis of the Tandem Beta-Zipper Interaction of a Streptococcal Protein with Human Fibronectin. Authors: Norris, N.C. / Bingham, R.J. / Harris, G. / Speakman, A. / Jones, R.P.O. / Leech, A. / Turkenburg, J.P. / Potts, J.R. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3zrz.cif.gz 3zrz.cif.gz | 61.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3zrz.ent.gz pdb3zrz.ent.gz | 44.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3zrz.json.gz 3zrz.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/zr/3zrz https://data.pdbj.org/pub/pdb/validation_reports/zr/3zrz ftp://data.pdbj.org/pub/pdb/validation_reports/zr/3zrz ftp://data.pdbj.org/pub/pdb/validation_reports/zr/3zrz | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2rkzS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 10120.389 Da / Num. of mol.: 2 / Fragment: SECOND AND THIRD F1 MODULES, RESIDUES 93-182 Source method: isolated from a genetically manipulated source Details: IN BOTH A AND B THERE ARE DISULFIDE BONDS BETWEEN RESIDUES C 97 AND C 125, C 123 AND C 135, C 141 AND C 169, C 167 AND C 179 Source: (gene. exp.)  HOMO SAPIENS (human) / Plasmid: PPIC9K / Production host: HOMO SAPIENS (human) / Plasmid: PPIC9K / Production host:  PICHIA PASTORIS (fungus) / Strain (production host): GS115 / References: UniProt: P02751 PICHIA PASTORIS (fungus) / Strain (production host): GS115 / References: UniProt: P02751#2: Protein/peptide | Mass: 1952.148 Da / Num. of mol.: 2 / Fragment: RESIDUES 560-577 / Source method: obtained synthetically Details: ACETYLATED ON THE N-TERMINUS AND AMIDATED ON THE C-TERMINUS Source: (synth.)  STREPTOCOCCUS PYOGENES (bacteria) / References: UniProt: Q01924, UniProt: Q711B0*PLUS STREPTOCOCCUS PYOGENES (bacteria) / References: UniProt: Q01924, UniProt: Q711B0*PLUS#3: Chemical | ChemComp-SO4 / | #4: Chemical | #5: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.45 Å3/Da / Density % sol: 49.7 % Description: 2F1 AND 3F1 FROM 2RKZ WERE SEARCHED SEPARATELY IN PHASER. THE PEPTIDE FROM THIS STRUCTURE WAS NOT USED. |
|---|---|
| Crystal grow | Temperature: 291 K / Method: vapor diffusion, sitting drop / pH: 7 Details: CRYSTALS OF THE PROTEIN/PEPTIDE COMPLEX (0.3 MM/3.5 MM, PH 7.7) WERE GROWN USING SITTING DROP VAPOUR DIFFUSION FROM A 1:1 DILUTION WITH WELL SOLUTION (0.1 M TRIS PH 7.0, 1.5 M (NH4)2SO4 AT 18 DEGREES CENTIGRADE |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID23-1 / Wavelength: 0.9804 / Beamline: ID23-1 / Wavelength: 0.9804 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Mar 8, 2008 / Details: BENT CYLINDRICAL MIRRORS |
| Radiation | Monochromator: SI(111) / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9804 Å / Relative weight: 1 |
| Reflection | Resolution: 1.7→28.22 Å / Num. obs: 25333 / % possible obs: 100 % / Observed criterion σ(I): 1.9 / Redundancy: 6.1 % / Biso Wilson estimate: 22.64 Å2 / Rmerge(I) obs: 0.05 / Net I/σ(I): 22.4 |
| Reflection shell | Resolution: 1.7→1.79 Å / Redundancy: 6.1 % / Rmerge(I) obs: 0.41 / Mean I/σ(I) obs: 3.6 / % possible all: 100 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2RKZ Resolution: 1.7→26.45 Å / Cor.coef. Fo:Fc: 0.964 / Cor.coef. Fo:Fc free: 0.938 / SU B: 2.041 / SU ML: 0.068 / Cross valid method: THROUGHOUT / ESU R: 0.102 / ESU R Free: 0.108 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS.
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 23.575 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.7→26.45 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller


























 PDBj
PDBj




















