+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3qor | ||||||
|---|---|---|---|---|---|---|---|
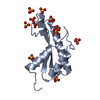
| Title | Crystal structure of human nuclear migration protein NudC | ||||||
 Components Components | (Nuclear migration protein ...) x 4 | ||||||
 Keywords Keywords | PROTEIN BINDING / CELL CYCLE / beta-sandwich / nuclear migration protein / chaperone | ||||||
| Function / homology |  Function and homology information Function and homology informationRND2 GTPase cycle / mitotic metaphase chromosome alignment / nuclear migration / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / Resolution of Sister Chromatid Cohesion / mitotic spindle organization / RHO GTPases Activate Formins / response to peptide hormone ...RND2 GTPase cycle / mitotic metaphase chromosome alignment / nuclear migration / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / Resolution of Sister Chromatid Cohesion / mitotic spindle organization / RHO GTPases Activate Formins / response to peptide hormone / spindle / mitotic spindle / : / Separation of Sister Chromatids / protein folding / midbody / microtubule / cadherin binding / cell division / nucleoplasm / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 1.753 Å SAD / Resolution: 1.753 Å | ||||||
 Authors Authors | Derewenda, U. / Derewenda, Z. / Zheng, M. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2011 Journal: J.Mol.Biol. / Year: 2011Title: Structural Features and Chaperone Activity of the NudC Protein Family. Authors: Zheng, M. / Cierpicki, T. / Burdette, A.J. / Utepbergenov, D. / Janczyk, P.L. / Derewenda, U. / Stukenberg, P.T. / Caldwell, K.A. / Derewenda, Z.S. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3qor.cif.gz 3qor.cif.gz | 151.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3qor.ent.gz pdb3qor.ent.gz | 119.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3qor.json.gz 3qor.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/qo/3qor https://data.pdbj.org/pub/pdb/validation_reports/qo/3qor ftp://data.pdbj.org/pub/pdb/validation_reports/qo/3qor ftp://data.pdbj.org/pub/pdb/validation_reports/qo/3qor | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| 3 | 
| ||||||||
| 4 | 
| ||||||||
| 5 | 
| ||||||||
| Unit cell |
|
- Components
Components
-Nuclear migration protein ... , 4 types, 5 molecules ABCDE
| #1: Protein | Mass: 13678.567 Da / Num. of mol.: 1 / Fragment: residues 158-274 / Mutation: E236A, K129A Source method: isolated from a genetically manipulated source Details: Cys188 is modified to S-OXY CYSTEINE / Source: (gene. exp.)  Homo sapiens (human) / Gene: NUDC / Production host: Homo sapiens (human) / Gene: NUDC / Production host:  |
|---|---|
| #2: Protein | Mass: 13694.566 Da / Num. of mol.: 1 / Fragment: residues 158-274 / Mutation: E236A, K129A Source method: isolated from a genetically manipulated source Details: Cys188 is modified to 3-SULFINOALANINE / Source: (gene. exp.)  Homo sapiens (human) / Gene: NUDC / Production host: Homo sapiens (human) / Gene: NUDC / Production host:  |
| #3: Protein | Mass: 13662.567 Da / Num. of mol.: 1 / Fragment: residues 158-274 / Mutation: E236A, K129A Source method: isolated from a genetically manipulated source Details: Cys188 is unmodified / Source: (gene. exp.)  Homo sapiens (human) / Gene: NUDC / Production host: Homo sapiens (human) / Gene: NUDC / Production host:  |
| #4: Protein | Mass: 13710.565 Da / Num. of mol.: 2 / Fragment: residues 158-274 / Mutation: E236A, K129A Source method: isolated from a genetically manipulated source Details: Cys188 is modified to CYSTEINESULFONIC ACID / Source: (gene. exp.)  Homo sapiens (human) / Gene: NUDC / Production host: Homo sapiens (human) / Gene: NUDC / Production host:  |
-Non-polymers , 2 types, 878 molecules 


| #5: Chemical | ChemComp-ACT / |
|---|---|
| #6: Water | ChemComp-HOH / |
-Details
| Has protein modification | Y |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.35 Å3/Da / Density % sol: 47.55 % |
|---|---|
| Crystal grow | Temperature: 296 K / Method: vapor diffusion, hanging drop / pH: 4.5 Details: 25% PEG 3350, 0.2M sodium acetate, 0.1M lithium sulfate, pH 4.5, VAPOR DIFFUSION, HANGING DROP, temperature 296K |
-Data collection
| Diffraction | Mean temperature: 110 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-ID / Wavelength: 0.97928 Å / Beamline: 22-ID / Wavelength: 0.97928 Å |
| Detector | Type: MAR scanner 300 mm plate / Detector: IMAGE PLATE / Date: May 5, 2005 |
| Radiation | Monochromator: Si 111 / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.97928 Å / Relative weight: 1 |
| Reflection | Resolution: 1.75→28.05 Å / Num. all: 63806 / Num. obs: 63750 / % possible obs: 99.9 % / Observed criterion σ(F): 1.35 / Observed criterion σ(I): 1 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD / Resolution: 1.753→28.05 Å / SU ML: 0.22 / σ(F): 1.35 / Stereochemistry target values: ML SAD / Resolution: 1.753→28.05 Å / SU ML: 0.22 / σ(F): 1.35 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.61 Å / VDW probe radii: 0.9 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 49.863 Å2 / ksol: 0.353 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.753→28.05 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
|
 Movie
Movie Controller
Controller












 PDBj
PDBj








