+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3nih | ||||||
|---|---|---|---|---|---|---|---|
| Title | The structure of UBR box (RIAAA) | ||||||
 Components Components |
| ||||||
 Keywords Keywords | METAL BINDING PROTEIN / E3 ubiquitin ligase / UBR box / Zinc-binding protein / N-end rule / Ligase | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of dipeptide transport / UBR1-RAD6 ubiquitin ligase complex / proteasome regulatory particle binding / stress-induced homeostatically regulated protein degradation pathway / ubiquitin-dependent protein catabolic process via the N-end rule pathway / mitochondria-associated ubiquitin-dependent protein catabolic process / cytoplasm protein quality control by the ubiquitin-proteasome system / ribosome-associated ubiquitin-dependent protein catabolic process / proteasome regulatory particle, base subcomplex / Antigen processing: Ubiquitination & Proteasome degradation ...regulation of dipeptide transport / UBR1-RAD6 ubiquitin ligase complex / proteasome regulatory particle binding / stress-induced homeostatically regulated protein degradation pathway / ubiquitin-dependent protein catabolic process via the N-end rule pathway / mitochondria-associated ubiquitin-dependent protein catabolic process / cytoplasm protein quality control by the ubiquitin-proteasome system / ribosome-associated ubiquitin-dependent protein catabolic process / proteasome regulatory particle, base subcomplex / Antigen processing: Ubiquitination & Proteasome degradation / cellular response to unfolded protein / protein monoubiquitination / ubiquitin ligase complex / ERAD pathway / RING-type E3 ubiquitin transferase / protein polyubiquitination / ubiquitin-protein transferase activity / ubiquitin protein ligase activity / protein ubiquitination / zinc ion binding / cytoplasm Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.1 Å MOLECULAR REPLACEMENT / Resolution: 2.1 Å | ||||||
 Authors Authors | Choi, W.S. / Jeong, B.-C. / Lee, M.-R. / Song, H.K. | ||||||
 Citation Citation |  Journal: Nat.Struct.Mol.Biol. / Year: 2010 Journal: Nat.Struct.Mol.Biol. / Year: 2010Title: Structural basis for the recognition of N-end rule substrates by the UBR box of ubiquitin ligases Authors: Choi, W.S. / Jeong, B.-C. / Joo, Y.J. / Lee, M.-R. / Kim, J. / Eck, M.J. / Song, H.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3nih.cif.gz 3nih.cif.gz | 31.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3nih.ent.gz pdb3nih.ent.gz | 20 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3nih.json.gz 3nih.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ni/3nih https://data.pdbj.org/pub/pdb/validation_reports/ni/3nih ftp://data.pdbj.org/pub/pdb/validation_reports/ni/3nih ftp://data.pdbj.org/pub/pdb/validation_reports/ni/3nih | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3niiC  3nijC  3nikC  3nilC  3nimC  3ninC  3nisSC  3nitC C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 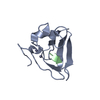
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 9236.394 Da / Num. of mol.: 1 / Fragment: UBR-type domain, residues 115-194 Source method: isolated from a genetically manipulated source Details: UBR box Source: (gene. exp.)  Plasmid: pET / Production host:  | ||||
|---|---|---|---|---|---|
| #2: Protein/peptide | Mass: 501.600 Da / Num. of mol.: 1 / Source method: obtained synthetically / Details: chemical synthesis | ||||
| #3: Chemical | | #4: Water | ChemComp-HOH / | Has protein modification | Y | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.79 Å3/Da / Density % sol: 55.97 % / Mosaicity: 0.696 ° |
|---|---|
| Crystal grow | Temperature: 295 K / Method: vapor diffusion, hanging drop / pH: 8 Details: 0.16M ammonium acetate, 0.01M calcium chloride dihydrate, 0.05M sodium cacodylate trihydrate pH 6.5, 8%(w/v) PEG 4000, VAPOR DIFFUSION, HANGING DROP, temperature 295K |
-Data collection
| Diffraction | Mean temperature: 100 K | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: PAL/PLS SYNCHROTRON / Site: PAL/PLS  / Beamline: 4A / Wavelength: 1 Å / Beamline: 4A / Wavelength: 1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Detector | Type: ADSC QUANTUM 210 / Detector: CCD / Date: Mar 25, 2009 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection | Resolution: 2.1→50 Å / Num. obs: 7049 / % possible obs: 99.8 % / Redundancy: 26.3 % / Rmerge(I) obs: 0.077 / Χ2: 2.217 / Net I/σ(I): 12.7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Reflection shell |
|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 3NIS Resolution: 2.1→37.35 Å / Cor.coef. Fo:Fc: 0.933 / Cor.coef. Fo:Fc free: 0.901 / Occupancy max: 1 / Occupancy min: 1 / SU B: 10.56 / SU ML: 0.172 / Cross valid method: THROUGHOUT / σ(F): 0 / ESU R Free: 0.192 / Stereochemistry target values: MAXIMUM LIKELIHOOD / Details: HYDROGENS HAVE BEEN ADDED IN THE RIDING POSITIONS
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Ion probe radii: 0.8 Å / Shrinkage radii: 0.8 Å / VDW probe radii: 1.2 Å / Solvent model: MASK | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso max: 78.5 Å2 / Biso mean: 59.135 Å2 / Biso min: 33.22 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.1→37.35 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.1→2.155 Å / Total num. of bins used: 20
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 18.1938 Å / Origin y: 9.7354 Å / Origin z: -6.8215 Å
|
 Movie
Movie Controller
Controller










 PDBj
PDBj






