[English] 日本語
 Yorodumi
Yorodumi- PDB-3if4: Structure from the mobile metagenome of North West Arm Sewage Out... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3if4 | ||||||
|---|---|---|---|---|---|---|---|
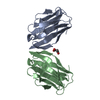
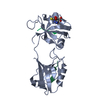
| Title | Structure from the mobile metagenome of North West Arm Sewage Outfall: Integron Cassette Protein Hfx_Cass5 | ||||||
 Components Components | Integron Cassette Protein Hfx_Cass5 | ||||||
 Keywords Keywords | structural genomics / unknown function / Integron Cassette Protein Mobile Metagenome Structural Genomics PSI-2 Protein Structure Initiative Midwest Center for Structural Genomics / MCSG | ||||||
| Function / homology | Integron cassette protein helical domain / Integron cassette protein / Anthopleurin-A / Single Sheet / Single alpha-helices involved in coiled-coils or other helix-helix interfaces / Up-down Bundle / Mainly Beta / Mainly Alpha Function and homology information Function and homology information | ||||||
| Biological species | unidentified (others) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 2.181 Å SAD / Resolution: 2.181 Å | ||||||
 Authors Authors | Sureshan, V. / Deshpande, C.N. / Harrop, S.J. / Evdokimova, E. / Kudrytska, M. / Koenig, J.E. / Osipiuk, J. / Edwards, A. / Savchenko, A. / Joachimiak, A. ...Sureshan, V. / Deshpande, C.N. / Harrop, S.J. / Evdokimova, E. / Kudrytska, M. / Koenig, J.E. / Osipiuk, J. / Edwards, A. / Savchenko, A. / Joachimiak, A. / Doolittle, W.F. / Stokes, H.W. / Curmi, P.M.G. / Mabbutt, B.C. / Midwest Center for Structural Genomics (MCSG) | ||||||
 Citation Citation |  Journal: Plos One / Year: 2013 Journal: Plos One / Year: 2013Title: Integron gene cassettes: a repository of novel protein folds with distinct interaction sites. Authors: Sureshan, V. / Deshpande, C.N. / Boucher, Y. / Koenig, J.E. / Stokes, H.W. / Harrop, S.J. / Curmi, P.M. / Mabbutt, B.C. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3if4.cif.gz 3if4.cif.gz | 174.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3if4.ent.gz pdb3if4.ent.gz | 140.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3if4.json.gz 3if4.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/if/3if4 https://data.pdbj.org/pub/pdb/validation_reports/if/3if4 ftp://data.pdbj.org/pub/pdb/validation_reports/if/3if4 ftp://data.pdbj.org/pub/pdb/validation_reports/if/3if4 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  3fuyC  3fxhC  3fy6C  3imoC  3jrtC C: citing same article ( |
|---|---|
| Similar structure data | |
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Details | The four chains in the asymmetric unit arrange as two dimers. Dimer 1: Chain A and Chain C Dimer 2: Chain B and Chain D |
- Components
Components
| #1: Protein | Mass: 13649.657 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Details: uncultured bacterium / Source: (gene. exp.) unidentified (others) / Plasmid: p15TV LIC / Production host:  #2: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.94 Å3/Da / Density % sol: 36.53 % |
|---|---|
| Crystal grow | Temperature: 294 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 29% PEG3350, 0.1 M Hepes pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 294K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 0.9798 Å / Beamline: 19-ID / Wavelength: 0.9798 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Feb 5, 2009 |
| Radiation | Monochromator: Si 111 / Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9798 Å / Relative weight: 1 |
| Reflection | Resolution: 2.18→50 Å / Num. obs: 21618 / % possible obs: 97.6 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 7 % / Biso Wilson estimate: 34.4 Å2 / Rmerge(I) obs: 0.093 / Rsym value: 0.093 / Net I/σ(I): 34.6 |
| Reflection shell | Resolution: 2.2→2.24 Å / Redundancy: 4 % / Rmerge(I) obs: 0.531 / Mean I/σ(I) obs: 2.62 / Rsym value: 0.531 / % possible all: 68.6 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD / Resolution: 2.181→40.226 Å / SU ML: 0.3 / Isotropic thermal model: anisotropic / σ(F): 0.05 / Phase error: 24.86 / Stereochemistry target values: ML SAD / Resolution: 2.181→40.226 Å / SU ML: 0.3 / Isotropic thermal model: anisotropic / σ(F): 0.05 / Phase error: 24.86 / Stereochemistry target values: ML
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 50.641 Å2 / ksol: 0.359 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.181→40.226 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Refine-ID: X-RAY DIFFRACTION
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: chain D and (resseq 53:98)) |
 Movie
Movie Controller
Controller










 PDBj
PDBj
