[English] 日本語
 Yorodumi
Yorodumi- PDB-3ge2: Crystal structure of putative lipoprotein SP_0198 from Streptococ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 3ge2 | ||||||
|---|---|---|---|---|---|---|---|
| Title | Crystal structure of putative lipoprotein SP_0198 from Streptococcus pneumoniae | ||||||
 Components Components | Lipoprotein, putative | ||||||
 Keywords Keywords | LIPOPROTEIN / beta-barrel / Structural Genomics / PSI-2 / Protein Structure Initiative / Midwest Center for Structural Genomics / MCSG | ||||||
| Function / homology |  Function and homology information Function and homology informationDomain of unknown function DUF3642, lipoprotein / Bacterial lipoprotein / Lipocalin - #50 / D-aminopeptidase/lipoprotein domain superfamily / Lipocalin / Prokaryotic membrane lipoprotein lipid attachment site profile. / Beta Barrel / Mainly Beta Similarity search - Domain/homology | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  SAD / Resolution: 2.203 Å SAD / Resolution: 2.203 Å | ||||||
 Authors Authors | Kim, Y. / Zhang, R. / Joachimiak, G. / Gornicki, P. / Joachimiak, A. / Midwest Center for Structural Genomics (MCSG) | ||||||
 Citation Citation |  Journal: To be Published Journal: To be PublishedTitle: Crystal Structure of Putative Lipoprotein SP_0198 from Streptococcus pneumoniae Authors: Kim, Y. / Zhang, R. / Joachimiak, G. / Gornicki, P. / Joachimiak, A. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  3ge2.cif.gz 3ge2.cif.gz | 52.4 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb3ge2.ent.gz pdb3ge2.ent.gz | 37 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  3ge2.json.gz 3ge2.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  3ge2_validation.pdf.gz 3ge2_validation.pdf.gz | 444 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  3ge2_full_validation.pdf.gz 3ge2_full_validation.pdf.gz | 446.1 KB | Display | |
| Data in XML |  3ge2_validation.xml.gz 3ge2_validation.xml.gz | 8.4 KB | Display | |
| Data in CIF |  3ge2_validation.cif.gz 3ge2_validation.cif.gz | 10.4 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ge/3ge2 https://data.pdbj.org/pub/pdb/validation_reports/ge/3ge2 ftp://data.pdbj.org/pub/pdb/validation_reports/ge/3ge2 ftp://data.pdbj.org/pub/pdb/validation_reports/ge/3ge2 | HTTPS FTP |
-Related structure data
| Similar structure data | |
|---|---|
| Other databases |
- Links
Links
- Assembly
Assembly
| Deposited unit | 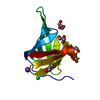
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| |||||||||
| Unit cell |
| |||||||||
| Components on special symmetry positions |
|
- Components
Components
| #1: Protein | Mass: 14113.791 Da / Num. of mol.: 1 / Fragment: UNP residues 26-152 Source method: isolated from a genetically manipulated source Details: N-terminal MBP-fusion with TVMV protease cut-site which is cleaved in vivo and a 6-His tag with TEV protease cut site Source: (gene. exp.)   | ||||||
|---|---|---|---|---|---|---|---|
| #2: Chemical | ChemComp-K / | ||||||
| #3: Chemical | ChemComp-CL / | ||||||
| #4: Chemical | ChemComp-GOL / #5: Water | ChemComp-HOH / | Has protein modification | Y | Sequence details | THE PROTEIN SEQUENCE PROVIDED BY AUTHORS IS THE SEQUENCE OF THE PROTEIN USED FOR CRYSTALLIZATION. ...THE PROTEIN SEQUENCE PROVIDED BY AUTHORS IS THE SEQUENCE OF THE PROTEIN USED FOR CRYSTALLIZ | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.71 Å3/Da / Density % sol: 66.87 % |
|---|---|
| Crystal grow | Temperature: 289 K / Method: vapor diffusion, sitting drop / pH: 4.5 Details: 0.1 M Sodium acetate trihydrate pH 4.5, 2.0 M Ammonium sulfate, VAPOR DIFFUSION, SITTING DROP, temperature 289K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 19-ID / Wavelength: 0.9793 Å / Beamline: 19-ID / Wavelength: 0.9793 Å |
| Detector | Type: ADSC QUANTUM 315 / Detector: CCD / Date: Feb 7, 2009 / Details: mirrors |
| Radiation | Monochromator: double crystal / Protocol: SAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9793 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→40.9 Å / Num. all: 11211 / Num. obs: 11211 / % possible obs: 99.1 % / Observed criterion σ(F): 0 / Observed criterion σ(I): 0 / Redundancy: 6.9 % / Biso Wilson estimate: 43.59 Å2 / Rmerge(I) obs: 0.078 / Net I/σ(I): 12.8 |
| Reflection shell | Resolution: 2.2→2.24 Å / Redundancy: 7.1 % / Rmerge(I) obs: 0.782 / Mean I/σ(I) obs: 3 / Num. unique all: 537 / % possible all: 98.7 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  SAD / Resolution: 2.203→37.421 Å / SU ML: 0.27 / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: MLHL SAD / Resolution: 2.203→37.421 Å / SU ML: 0.27 / Cross valid method: THROUGHOUT / σ(F): 0 / Stereochemistry target values: MLHL
| ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 74.727 Å2 / ksol: 0.399 e/Å3 | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters |
| ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.203→37.421 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Refine-ID: X-RAY DIFFRACTION / Total num. of bins used: 4
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Origin x: 25.2015 Å / Origin y: 31.4046 Å / Origin z: 32.0837 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group | Selection details: chain A |
 Movie
Movie Controller
Controller












 PDBj
PDBj











