| ソフトウェア | | 名称 | バージョン | 分類 |
|---|
| XDS | | データスケーリング | | AUTOMAR | | データ削減 | | AMoRE | | 位相決定 |  X-PLOR X-PLOR | 3.851 | 精密化 | | XDS | | データ削減 |
|
|---|
| 精密化 | 構造決定の手法:  分子置換 分子置換
開始モデル: P61 OCTAMER
解像度: 2.15→7 Å / σ(F): 2 / | Rfactor | 反射数 |
|---|
| Rwork | 0.166 | - |
|---|
| obs | 0.1661 | 3827 |
|---|
| all | - | 4238 |
|---|
|
|---|
| 精密化ステップ | サイクル: LAST / 解像度: 2.15→7 Å
| タンパク質 | 核酸 | リガンド | 溶媒 | 全体 |
|---|
| 原子数 | 0 | 404 | 13 | 82 | 499 |
|---|
|
|---|
| 拘束条件 | | Refine-ID | タイプ | Dev ideal |
|---|
| X-RAY DIFFRACTION | x_bond_d| 0.011 | | X-RAY DIFFRACTION | x_bond_d_na | | X-RAY DIFFRACTION | x_bond_d_prot | | X-RAY DIFFRACTION | x_angle_d | | X-RAY DIFFRACTION | x_angle_d_na | | X-RAY DIFFRACTION | x_angle_d_prot | | X-RAY DIFFRACTION | x_angle_deg| 1.4 | | X-RAY DIFFRACTION | x_angle_deg_na | | X-RAY DIFFRACTION | x_angle_deg_prot | | X-RAY DIFFRACTION | x_dihedral_angle_d| 27 | | X-RAY DIFFRACTION | x_dihedral_angle_d_na | | X-RAY DIFFRACTION | x_dihedral_angle_d_prot | | X-RAY DIFFRACTION | x_improper_angle_d| 1.6 | | X-RAY DIFFRACTION | x_improper_angle_d_na | | X-RAY DIFFRACTION | x_improper_angle_d_prot | | X-RAY DIFFRACTION | x_mcbond_it | | X-RAY DIFFRACTION | x_mcangle_it | | X-RAY DIFFRACTION | x_scbond_it | | X-RAY DIFFRACTION | x_scangle_it | | | | | | | | | | | | | | | | | | | |
|
|---|
| Xplor file | Serial no: 1 / Param file: DNA.PARAM / Topol file: DNA.TOP |
|---|
| ソフトウェア | *PLUS 名称:  X-PLOR / バージョン: 3.851 / 分類: refinement X-PLOR / バージョン: 3.851 / 分類: refinement |
|---|
| 精密化 | *PLUS 最高解像度: 2.15 Å / 最低解像度: 7 Å |
|---|
| 溶媒の処理 | *PLUS |
|---|
| 原子変位パラメータ | *PLUS |
|---|
| 拘束条件 | *PLUS | Refine-ID | タイプ | Dev ideal |
|---|
| X-RAY DIFFRACTION | x_dihedral_angle_d | | X-RAY DIFFRACTION | x_dihedral_angle_deg| 27 | | X-RAY DIFFRACTION | x_improper_angle_d | | X-RAY DIFFRACTION | x_improper_angle_deg| 1.6 | | | | |
|
|---|
 データを開く
データを開く 基本情報
基本情報 要素
要素 キーワード
キーワード 機能・相同性情報
機能・相同性情報 X線回折 /
X線回折 /  シンクロトロン /
シンクロトロン /  分子置換 / 解像度: 2.15 Å
分子置換 / 解像度: 2.15 Å  データ登録者
データ登録者 引用
引用 ジャーナル: Embo J. / 年: 1998
ジャーナル: Embo J. / 年: 1998 構造の表示
構造の表示 Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク ダウンロード
ダウンロード 384d.cif.gz
384d.cif.gz PDBx/mmCIF形式
PDBx/mmCIF形式 pdb384d.ent.gz
pdb384d.ent.gz PDB形式
PDB形式 384d.json.gz
384d.json.gz PDBx/mmJSON形式
PDBx/mmJSON形式 その他のダウンロード
その他のダウンロード https://data.pdbj.org/pub/pdb/validation_reports/84/384d
https://data.pdbj.org/pub/pdb/validation_reports/84/384d ftp://data.pdbj.org/pub/pdb/validation_reports/84/384d
ftp://data.pdbj.org/pub/pdb/validation_reports/84/384d リンク
リンク 集合体
集合体
 要素
要素 X線回折 / 使用した結晶の数: 1
X線回折 / 使用した結晶の数: 1  試料調製
試料調製 シンクロトロン / サイト: LURE
シンクロトロン / サイト: LURE  / ビームライン: DW32
/ ビームライン: DW32 解析
解析 分子置換
分子置換 X-PLOR / バージョン: 3.851 / 分類: refinement
X-PLOR / バージョン: 3.851 / 分類: refinement ムービー
ムービー コントローラー
コントローラー









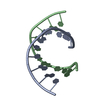


 PDBj
PDBj




