[English] 日本語
 Yorodumi
Yorodumi- PDB-2z7r: Crystal Structure of the N-terminal Kinase Domain of Human RSK1 b... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2z7r | ||||||
|---|---|---|---|---|---|---|---|
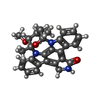
| Title | Crystal Structure of the N-terminal Kinase Domain of Human RSK1 bound to Staurosporine | ||||||
 Components Components | Ribosomal protein S6 kinase alpha-1 | ||||||
 Keywords Keywords | TRANSFERASE / protein kinase / cancer / kinase inhibitor / ATP-binding / Magnesium / Metal-binding / Nucleotide-binding / Phosphorylation / Polymorphism / Serine/threonine-protein kinase | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of translation in response to stress / CREB1 phosphorylation through NMDA receptor-mediated activation of RAS signaling / ribosomal protein S6 kinase activity / hepatocyte proliferation / CREB phosphorylation / positive regulation of hepatic stellate cell activation / Gastrin-CREB signalling pathway via PKC and MAPK / RSK activation / negative regulation of TOR signaling / ERK/MAPK targets ...regulation of translation in response to stress / CREB1 phosphorylation through NMDA receptor-mediated activation of RAS signaling / ribosomal protein S6 kinase activity / hepatocyte proliferation / CREB phosphorylation / positive regulation of hepatic stellate cell activation / Gastrin-CREB signalling pathway via PKC and MAPK / RSK activation / negative regulation of TOR signaling / ERK/MAPK targets / Recycling pathway of L1 / TORC1 signaling / protein serine/threonine/tyrosine kinase activity / Transcriptional and post-translational regulation of MITF-M expression and activity / positive regulation of cell differentiation / positive regulation of cell growth / Senescence-Associated Secretory Phenotype (SASP) / chemical synaptic transmission / protein phosphorylation / non-specific serine/threonine protein kinase / protein serine kinase activity / protein serine/threonine kinase activity / regulation of DNA-templated transcription / synapse / negative regulation of apoptotic process / positive regulation of DNA-templated transcription / magnesium ion binding / signal transduction / positive regulation of transcription by RNA polymerase II / nucleoplasm / ATP binding / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Ikuta, M. / Munshi, S.K. | ||||||
 Citation Citation |  Journal: Protein Sci. / Year: 2007 Journal: Protein Sci. / Year: 2007Title: Crystal structures of the N-terminal kinase domain of human RSK1 bound to three different ligands: Implications for the design of RSK1 specific inhibitors. Authors: Ikuta, M. / Kornienko, M. / Byrne, N. / Reid, J.C. / Mizuarai, S. / Kotani, H. / Munshi, S.K. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2z7r.cif.gz 2z7r.cif.gz | 71.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2z7r.ent.gz pdb2z7r.ent.gz | 51.4 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2z7r.json.gz 2z7r.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/z7/2z7r https://data.pdbj.org/pub/pdb/validation_reports/z7/2z7r ftp://data.pdbj.org/pub/pdb/validation_reports/z7/2z7r ftp://data.pdbj.org/pub/pdb/validation_reports/z7/2z7r | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2z7qC  2z7sC  1vzoS C: citing same article ( S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 36777.391 Da / Num. of mol.: 1 / Fragment: Residues 33-353 Source method: isolated from a genetically manipulated source Source: (gene. exp.)  Homo sapiens (human) / Gene: RPS6KA1, RSK1 / Cell line (production host): SF21 / Production host: Homo sapiens (human) / Gene: RPS6KA1, RSK1 / Cell line (production host): SF21 / Production host:  References: UniProt: Q15418, non-specific serine/threonine protein kinase |
|---|---|
| #2: Chemical | ChemComp-STU / |
| #3: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.3 Å3/Da / Density % sol: 46.41 % |
|---|---|
| Crystal grow | Temperature: 293 K / Method: vapor diffusion, hanging drop / pH: 7.5 Details: 12-16% PEG MME 2000, 150mM DL-malic acid, pH 7.5, VAPOR DIFFUSION, HANGING DROP, temperature 293K |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 17-ID / Beamline: 17-ID |
| Detector | Type: ADSC QUANTUM 210 / Detector: CCD / Date: Jul 27, 2006 |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Relative weight: 1 |
| Reflection | Resolution: 2→50 Å / Num. obs: 23744 / % possible obs: 98.6 % / Redundancy: 6 % / Biso Wilson estimate: 31.2 Å2 |
- Processing
Processing
| Software |
| ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: 1VZO Resolution: 2→20 Å
| ||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→20 Å
|
 Movie
Movie Controller
Controller












 PDBj
PDBj