[English] 日本語
 Yorodumi
Yorodumi- PDB-2xc9: BINARY COMPLEX OF SULFOLOBUS SOLFATARICUS DPO4 DNA POLYMERASE AND... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2xc9 | ||||||
|---|---|---|---|---|---|---|---|
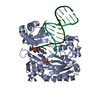
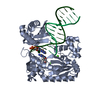
| Title | BINARY COMPLEX OF SULFOLOBUS SOLFATARICUS DPO4 DNA POLYMERASE AND 1, N2-ETHENOGUANINE MODIFIED DNA, MAGNESIUM FORM | ||||||
 Components Components |
| ||||||
 Keywords Keywords | TRANSFERASE/DNA / TRANSFERASE-DNA COMPLEX / TRANSLESION NUCLEOTIDYLTRANSFERASE | ||||||
| Function / homology |  Function and homology information Function and homology informationerror-prone translesion synthesis / DNA-templated DNA replication / DNA-directed DNA polymerase / damaged DNA binding / DNA-directed DNA polymerase activity / DNA repair / magnesium ion binding / cytoplasm Similarity search - Function | ||||||
| Biological species |   SULFOLOBUS SOLFATARICUS (archaea) SULFOLOBUS SOLFATARICUS (archaea) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.2 Å MOLECULAR REPLACEMENT / Resolution: 2.2 Å | ||||||
 Authors Authors | Irimia, A. / Loukachevitch, L.V. / Egli, M. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.F / Year: 2010 Journal: Acta Crystallogr.,Sect.F / Year: 2010Title: Metal Ion Dependence of the Active Site Conformation of the Trans-Lesion DNA Polymerase Dpo4 from Sulfolobus Solfataricus Authors: Irimia, A. / Loukachevitch, L.V. / Eoff, R.L. / Guengerich, F.P. / Egli, M. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2xc9.cif.gz 2xc9.cif.gz | 109.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2xc9.ent.gz pdb2xc9.ent.gz | 80.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2xc9.json.gz 2xc9.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/xc/2xc9 https://data.pdbj.org/pub/pdb/validation_reports/xc/2xc9 ftp://data.pdbj.org/pub/pdb/validation_reports/xc/2xc9 ftp://data.pdbj.org/pub/pdb/validation_reports/xc/2xc9 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2xcaC  2xcpC  2bq3S S: Starting model for refinement C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 41086.754 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   SULFOLOBUS SOLFATARICUS (archaea) / Strain: P2 / Gene: DPO4 / Plasmid: PET22B/DPO4-NHIS / Production host: SULFOLOBUS SOLFATARICUS (archaea) / Strain: P2 / Gene: DPO4 / Plasmid: PET22B/DPO4-NHIS / Production host:  |
|---|---|
| #2: DNA chain | Mass: 4425.876 Da / Num. of mol.: 1 / Source method: obtained synthetically |
| #3: DNA chain | Mass: 5396.503 Da / Num. of mol.: 1 / Source method: obtained synthetically |
| #4: Water | ChemComp-HOH / |
| Sequence details | THE N-TERMINAL 6 RESIDUES HIS TAG ARE DISORDERED AND MISSING FROM THE STRUCTURE. THE C-TERMINAL 11 ...THE N-TERMINAL 6 RESIDUES HIS TAG ARE DISORDERED |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 3.16 Å3/Da / Density % sol: 48.08 % Description: THE MODEL POSITION WAS OPTIMIZED BY SEVERAL ROUNDS OF RIGID BODY REFINEMENT. |
|---|---|
| Crystal grow | pH: 7.5 Details: 12-20% POLYETHYLENE GLYCOL 3350, 0.2 M AMMONIUM ACETATE, 0.1 M MAGNESIUM ACETATE, AND 20 MM TRIS (PH 7.5) |
-Data collection
| Diffraction | Mean temperature: 110 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  APS APS  / Beamline: 22-BM / Wavelength: 1 / Beamline: 22-BM / Wavelength: 1 |
| Detector | Type: MARRESEARCH / Detector: CCD / Date: Mar 12, 2006 Details: ROSENBAUM-ROCK MONOCHROMATOR HIGH -RESOLUTION DOUBLE-CRYSTAL SI (111) SAGITTAL FOCUSING. ROSENBAUM-ROCK VERTICAL FOCUSING MIRROR |
| Radiation | Protocol: SINGLE WAVELENGTH / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2.2→26.91 Å / Num. obs: 26245 / % possible obs: 99.3 % / Observed criterion σ(I): 1 / Redundancy: 12.6 % / Biso Wilson estimate: 43.4 Å2 / Rmerge(I) obs: 0.06 / Net I/σ(I): 36.3 |
| Reflection shell | Resolution: 2.2→2.28 Å / Redundancy: 10.4 % / Rmerge(I) obs: 0.48 / Mean I/σ(I) obs: 6.2 / % possible all: 99.3 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2BQ3 Resolution: 2.2→26.91 Å / Rfactor Rfree error: 0.007 / Data cutoff high absF: 1362679.09 / Data cutoff low absF: 0 / Isotropic thermal model: RESTRAINED / Cross valid method: THROUGHOUT / σ(F): 0
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Solvent model: FLAT MODEL / Bsol: 42.325 Å2 / ksol: 0.309453 e/Å3 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 40.7 Å2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.2→26.91 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints NCS | NCS model details: NONE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.2→2.34 Å / Rfactor Rfree error: 0.02 / Total num. of bins used: 6
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
|
 Movie
Movie Controller
Controller

































































 PDBj
PDBj







































