[English] 日本語
 Yorodumi
Yorodumi- PDB-2x40: Structure of beta-glucosidase 3B from Thermotoga neapolitana in c... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2x40 | ||||||
|---|---|---|---|---|---|---|---|
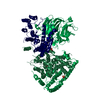
| Title | Structure of beta-glucosidase 3B from Thermotoga neapolitana in complex with glycerol | ||||||
 Components Components | BETA-GLUCOSIDASE | ||||||
 Keywords Keywords | HYDROLASE / TIM BARREL FOLD / FIBRONECTIN TYPE III FOLD | ||||||
| Function / homology |  Function and homology information Function and homology informationbeta-glucosidase / beta-glucosidase activity / carbohydrate metabolic process Similarity search - Function | ||||||
| Biological species |   THERMOTOGA NEAPOLITANA (bacteria) THERMOTOGA NEAPOLITANA (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 2.313 Å MAD / Resolution: 2.313 Å | ||||||
 Authors Authors | Pozzo, T. / Karlsson, E.N. / Logan, D.T. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 2010 Journal: J.Mol.Biol. / Year: 2010Title: Structural and Functional Analysis of Beta-Glucosidase 3B from Thermotoga Neapolitana: A Thermostable 3-Domain Representative of Glycoside Hydrolase Family 3 Authors: Pozzo, T. / Linares Pasten, J. / Karlsson, E.N. / Logan, D.T. #1: Journal: Acta Crystallogr.,Sect.F / Year: 2007 Title: Expression, Purification, Crystallization and Preliminary X-Ray Diffraction Analysis of Thermotoga Neapolitana Beta-Glucosidase B. Authors: Turner, P. / Pramhed, A. / Kanders, E. / Hedstrom, M. / Karlsson, E.N. / Logan, D.T. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2x40.cif.gz 2x40.cif.gz | 296.3 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2x40.ent.gz pdb2x40.ent.gz | 243 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2x40.json.gz 2x40.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/x4/2x40 https://data.pdbj.org/pub/pdb/validation_reports/x4/2x40 ftp://data.pdbj.org/pub/pdb/validation_reports/x4/2x40 ftp://data.pdbj.org/pub/pdb/validation_reports/x4/2x40 | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2x41C  2x42C  2wt3  2wt5  2wt6 C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 81261.867 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   THERMOTOGA NEAPOLITANA (bacteria) / Strain: DSM 4359 / Description: GERMAN COLLECTION OF MICROORGANISMS (DSM) / Production host: THERMOTOGA NEAPOLITANA (bacteria) / Strain: DSM 4359 / Description: GERMAN COLLECTION OF MICROORGANISMS (DSM) / Production host:  | ||||
|---|---|---|---|---|---|
| #2: Chemical | ChemComp-BR / #3: Chemical | ChemComp-GOL / | #4: Water | ChemComp-HOH / | |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.57 Å3/Da / Density % sol: 57 % / Description: NONE |
|---|---|
| Crystal grow | pH: 7.4 Details: PROTEIN 3-5 MG/ML, 16% PEG 3350, 0.1 M KBR, 90 MM BIS-TRIS PROPANE PH 7.4 |
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  ESRF ESRF  / Beamline: ID29 / Wavelength: 0.9791 / Beamline: ID29 / Wavelength: 0.9791 |
| Detector | Type: ADSC CCD / Detector: CCD / Date: Sep 21, 2006 / Details: MULTILAYER MIRRORS |
| Radiation | Protocol: MAD / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 0.9791 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→34.5 Å / Num. obs: 37161 / % possible obs: 98.8 % / Observed criterion σ(I): 0 / Redundancy: 4.5 % / Biso Wilson estimate: 43.22 Å2 / Rmerge(I) obs: 0.08 / Net I/σ(I): 13.2 |
| Reflection shell | Resolution: 2.3→2.4 Å / Rmerge(I) obs: 0.52 / Mean I/σ(I) obs: 2.3 / % possible all: 93.4 |
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MAD MADStarting model: NONE Resolution: 2.313→34.453 Å / SU ML: 0.32 / σ(F): 1.37 / Phase error: 25.34 / Stereochemistry target values: ML
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | Shrinkage radii: 0.9 Å / VDW probe radii: 1.11 Å / Solvent model: FLAT BULK SOLVENT MODEL / Bsol: 50.019 Å2 / ksol: 0.355 e/Å3 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 55.5 Å2
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.313→34.453 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS params. | Method: refined / Refine-ID: X-RAY DIFFRACTION
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement TLS group |
|
 Movie
Movie Controller
Controller











 PDBj
PDBj






