| Entry | Database: PDB / ID: 2wbs
|
|---|
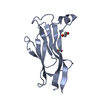
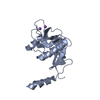
| Title | Crystal structure of the zinc finger domain of Klf4 bound to its target DNA |
|---|
 Components Components | - 5'-D(*GP*AP*GP*GP*CP*GP*CP)-3'
- 5'-D(*GP*CP*GP*CP*CP*TP*CP)-3'
- KRUEPPEL-LIKE FACTOR 4
|
|---|
 Keywords Keywords | TRANSCRIPTION/DNA / TRANSCRIPTION-DNA COMPLEX / DNA-BINDING / TRANSCRIPTION / METAL-BINDING / DNA / PROTEIN / NUCLEUS / ACTIVATOR / ZINC-FINGER / TRANSCRIPTION REGULATION |
|---|
| Function / homology |  Function and homology information Function and homology information
negative regulation of leukocyte adhesion to arterial endothelial cell / regulation of blastocyst development / : / negative regulation of chemokine (C-X-C motif) ligand 2 production / cellular response to endothelin / post-embryonic camera-type eye development / positive regulation of hemoglobin biosynthetic process / negative regulation of response to cytokine stimulus / epidermal cell differentiation / negative regulation of heterotypic cell-cell adhesion ...negative regulation of leukocyte adhesion to arterial endothelial cell / regulation of blastocyst development / : / negative regulation of chemokine (C-X-C motif) ligand 2 production / cellular response to endothelin / post-embryonic camera-type eye development / positive regulation of hemoglobin biosynthetic process / negative regulation of response to cytokine stimulus / epidermal cell differentiation / negative regulation of heterotypic cell-cell adhesion / negative regulation of muscle hyperplasia / epidermis morphogenesis / cellular response to laminar fluid shear stress / phosphatidylinositol 3-kinase regulator activity / negative regulation of interleukin-8 production / regulation of axon regeneration / negative regulation of cell migration involved in sprouting angiogenesis / post-embryonic hemopoiesis / defense response to tumor cell / negative regulation of G1/S transition of mitotic cell cycle / stem cell population maintenance / positive regulation of sprouting angiogenesis / lncRNA binding / positive regulation of telomere maintenance / positive regulation of protein metabolic process / regulation of cell differentiation / establishment of skin barrier / epidermis development / negative regulation of extrinsic apoptotic signaling pathway in absence of ligand / response to retinoic acid / canonical Wnt signaling pathway / fat cell differentiation / cellular response to retinoic acid / negative regulation of angiogenesis / negative regulation of cell migration / negative regulation of phosphatidylinositol 3-kinase/protein kinase B signal transduction / transcription coregulator binding / cellular response to leukemia inhibitory factor / negative regulation of canonical NF-kappaB signal transduction / negative regulation of smooth muscle cell proliferation / promoter-specific chromatin binding / euchromatin / chromatin DNA binding / negative regulation of ERK1 and ERK2 cascade / positive regulation of miRNA transcription / beta-catenin binding / cellular response to growth factor stimulus / histone deacetylase binding / cellular response to hydrogen peroxide / positive regulation of nitric oxide biosynthetic process / sequence-specific double-stranded DNA binding / regulation of cell population proliferation / microtubule cytoskeleton / double-stranded DNA binding / DNA-binding transcription activator activity, RNA polymerase II-specific / transcription regulator complex / sequence-specific DNA binding / gene expression / RNA polymerase II-specific DNA-binding transcription factor binding / DNA-binding transcription factor activity, RNA polymerase II-specific / cell differentiation / transcription cis-regulatory region binding / RNA polymerase II cis-regulatory region sequence-specific DNA binding / DNA-binding transcription factor activity / negative regulation of cell population proliferation / negative regulation of gene expression / negative regulation of DNA-templated transcription / positive regulation of gene expression / regulation of transcription by RNA polymerase II / DNA-templated transcription / positive regulation of DNA-templated transcription / chromatin / negative regulation of transcription by RNA polymerase II / positive regulation of transcription by RNA polymerase II / DNA binding / zinc ion binding / nucleoplasm / nucleus / cytoplasm / cytosolSimilarity search - Function Classic Zinc Finger / Double Stranded RNA Binding Domain / Zinc finger, C2H2 type / zinc finger / Zinc finger C2H2 type domain profile. / Zinc finger C2H2 superfamily / Zinc finger C2H2 type domain signature. / Zinc finger C2H2-type / 2-Layer Sandwich / Alpha BetaSimilarity search - Domain/homology |
|---|
| Biological species |   MUS MUSCULUS (house mouse) MUS MUSCULUS (house mouse) |
|---|
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MAD / Resolution: 1.7 Å MAD / Resolution: 1.7 Å |
|---|
 Authors Authors | Zocher, G. / Schuetz, A. / Carstanjen, D. / Heinemann, U. |
|---|
 Citation Citation |  Journal: Cell.Mol.Life Sci. / Year: 2011 Journal: Cell.Mol.Life Sci. / Year: 2011
Title: The Structure of the Klf4 DNA-Binding Domain Links to Self-Renewal and Macrophage Differentiation.
Authors: Schuetz, A. / Nana, D. / Rose, C. / Zocher, G. / Milanovic, M. / Koenigsmann, J. / Blasig, R. / Heinemann, U. / Carstanjen, D. |
|---|
| History | | Deposition | Mar 3, 2009 | Deposition site: PDBE / Processing site: PDBE |
|---|
| Revision 1.0 | Apr 7, 2010 | Provider: repository / Type: Initial release |
|---|
| Revision 1.1 | Jul 13, 2011 | Group: Advisory / Version format compliance |
|---|
| Revision 1.2 | Oct 5, 2011 | Group: Database references / Source and taxonomy |
|---|
| Revision 1.3 | May 8, 2024 | Group: Data collection / Database references ...Data collection / Database references / Derived calculations / Other
Category: chem_comp_atom / chem_comp_bond ...chem_comp_atom / chem_comp_bond / database_2 / pdbx_database_status / pdbx_struct_conn_angle / struct_conn / struct_site
Item: _database_2.pdbx_DOI / _database_2.pdbx_database_accession ..._database_2.pdbx_DOI / _database_2.pdbx_database_accession / _pdbx_database_status.status_code_sf / _pdbx_struct_conn_angle.ptnr1_auth_comp_id / _pdbx_struct_conn_angle.ptnr1_auth_seq_id / _pdbx_struct_conn_angle.ptnr1_label_atom_id / _pdbx_struct_conn_angle.ptnr1_label_comp_id / _pdbx_struct_conn_angle.ptnr1_label_seq_id / _pdbx_struct_conn_angle.ptnr2_auth_seq_id / _pdbx_struct_conn_angle.ptnr2_label_asym_id / _pdbx_struct_conn_angle.ptnr3_auth_comp_id / _pdbx_struct_conn_angle.ptnr3_auth_seq_id / _pdbx_struct_conn_angle.ptnr3_label_atom_id / _pdbx_struct_conn_angle.ptnr3_label_comp_id / _pdbx_struct_conn_angle.ptnr3_label_seq_id / _pdbx_struct_conn_angle.value / _struct_conn.pdbx_dist_value / _struct_conn.ptnr1_auth_comp_id / _struct_conn.ptnr1_auth_seq_id / _struct_conn.ptnr1_label_asym_id / _struct_conn.ptnr1_label_atom_id / _struct_conn.ptnr1_label_comp_id / _struct_conn.ptnr1_label_seq_id / _struct_conn.ptnr2_auth_comp_id / _struct_conn.ptnr2_auth_seq_id / _struct_conn.ptnr2_label_asym_id / _struct_conn.ptnr2_label_atom_id / _struct_conn.ptnr2_label_comp_id / _struct_conn.ptnr2_label_seq_id / _struct_site.pdbx_auth_asym_id / _struct_site.pdbx_auth_comp_id / _struct_site.pdbx_auth_seq_id |
|---|
|
|---|
 Yorodumi
Yorodumi Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MAD / Resolution: 1.7 Å
MAD / Resolution: 1.7 Å  Authors
Authors Citation
Citation Journal: Cell.Mol.Life Sci. / Year: 2011
Journal: Cell.Mol.Life Sci. / Year: 2011 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 2wbs.cif.gz
2wbs.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb2wbs.ent.gz
pdb2wbs.ent.gz PDB format
PDB format 2wbs.json.gz
2wbs.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/wb/2wbs
https://data.pdbj.org/pub/pdb/validation_reports/wb/2wbs ftp://data.pdbj.org/pub/pdb/validation_reports/wb/2wbs
ftp://data.pdbj.org/pub/pdb/validation_reports/wb/2wbs Links
Links Assembly
Assembly
 Components
Components








 X-RAY DIFFRACTION / Number of used crystals: 2
X-RAY DIFFRACTION / Number of used crystals: 2  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  BESSY
BESSY  / Beamline: 14.2 / Wavelength: 0.91841, 2.0, 1.28204, 1.28312, 1.16967
/ Beamline: 14.2 / Wavelength: 0.91841, 2.0, 1.28204, 1.28312, 1.16967 Processing
Processing MAD
MAD Movie
Movie Controller
Controller











 PDBj
PDBj







