[English] 日本語
 Yorodumi
Yorodumi- PDB-2tmv: VISUALIZATION OF PROTEIN-NUCLEIC ACID INTERACTIONS IN A VIRUS. RE... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 2tmv | ||||||
|---|---|---|---|---|---|---|---|
| Title | VISUALIZATION OF PROTEIN-NUCLEIC ACID INTERACTIONS IN A VIRUS. REFINED STRUCTURE OF INTACT TOBACCO MOSAIC VIRUS AT 2.9 ANGSTROMS RESOLUTION BY X-RAY FIBER DIFFRACTION | ||||||
 Components Components |
| ||||||
 Keywords Keywords | Virus/RNA / VIRUS / Helical virus / Virus-RNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology informationhelical viral capsid / structural molecule activity / identical protein binding Similarity search - Function | ||||||
| Biological species |   Tobacco mosaic virus Tobacco mosaic virus | ||||||
| Method |  FIBER DIFFRACTION / Resolution: 2.9 Å FIBER DIFFRACTION / Resolution: 2.9 Å | ||||||
 Authors Authors | Stubbs, G. / Pattanayek, R. / Namba, K. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1989 Journal: J.Mol.Biol. / Year: 1989Title: Visualization of protein-nucleic acid interactions in a virus. Refined structure of intact tobacco mosaic virus at 2.9 A resolution by X-ray fiber diffraction. Authors: Namba, K. / Pattanayek, R. / Stubbs, G. #1:  Journal: Science / Year: 1986 Journal: Science / Year: 1986Title: Structure of Tobacco Mosaic Virus at 3.6 Angstroms Resolution. Implications for Assembly Authors: Namba, K. / Stubbs, G. #2:  Journal: Biophys.J. / Year: 1986 Journal: Biophys.J. / Year: 1986Title: Application of Restrained Least-Squares Refinement to Fiber Diffraction from Macromolecular Assemblies Authors: Stubbs, G. / Namba, K. / Makowski, L. #3:  Journal: Acta Crystallogr.,Sect.A / Year: 1985 Journal: Acta Crystallogr.,Sect.A / Year: 1985Title: Solving the Phase Problem in Fiber Diffraction. Application to Tobacco Mosaic Virus at 3.6 Angstroms Resolution Authors: Namba, K. / Stubbs, G. #4:  Journal: J.Mol.Biol. / Year: 1981 Journal: J.Mol.Biol. / Year: 1981Title: Structure of the RNA in Tobacco Mosaic Virus Authors: Stubbs, G. / Stauffacher, C. #5:  Journal: Nature / Year: 1977 Journal: Nature / Year: 1977Title: Structure of RNA and RNA Binding Site in Tobacco Mosaic Virus from 4-Angstroms Map Calculated from X-Ray Fibre Diagrams Authors: Stubbs, G. / Warren, S. / Holmes, K. | ||||||
| History |
| ||||||
| Remark 650 | HELIX THE ENDS OF ALPHA HELICES ARE OFTEN DIFFICULT TO DEFINE AND IN SEVERAL CASES INCLUDE ONE TURN ...HELIX THE ENDS OF ALPHA HELICES ARE OFTEN DIFFICULT TO DEFINE AND IN SEVERAL CASES INCLUDE ONE TURN OF 3/10 HELIX. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  2tmv.cif.gz 2tmv.cif.gz | 49.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb2tmv.ent.gz pdb2tmv.ent.gz | 34.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  2tmv.json.gz 2tmv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Summary document |  2tmv_validation.pdf.gz 2tmv_validation.pdf.gz | 363.5 KB | Display |  wwPDB validaton report wwPDB validaton report |
|---|---|---|---|---|
| Full document |  2tmv_full_validation.pdf.gz 2tmv_full_validation.pdf.gz | 397.3 KB | Display | |
| Data in XML |  2tmv_validation.xml.gz 2tmv_validation.xml.gz | 10.1 KB | Display | |
| Data in CIF |  2tmv_validation.cif.gz 2tmv_validation.cif.gz | 13.8 KB | Display | |
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/tm/2tmv https://data.pdbj.org/pub/pdb/validation_reports/tm/2tmv ftp://data.pdbj.org/pub/pdb/validation_reports/tm/2tmv ftp://data.pdbj.org/pub/pdb/validation_reports/tm/2tmv | HTTPS FTP |
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | x 49
| ||||||||
| 2 |
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: SEE REMARK 7. | ||||||||
| Symmetry | Helical symmetry: (Circular symmetry: 1 / Dyad axis: no / N subunits divisor: 49 / Num. of operations: 49 / Rise per n subunits: 69 Å / Rotation per n subunits: 1080 °) | ||||||||
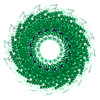
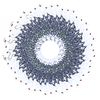
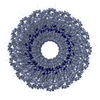
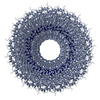
| Details | TMV IS A ROD-SHAPED VIRUS 3000 ANGSTROMS LONG AND 180 ANGSTROMS IN DIAMETER, WITH A CENTRAL HOLE OF DIAMETER 40 ANGSTROMS. APPROXIMATELY 2130 IDENTICAL PROTEIN SUBUNITS OF MOLECULAR WEIGHT 17500 FORM A RIGHT-HANDED HELIX OF PITCH 23 ANGSTROMS AND LENGTH 69 ANGTROMS WITH 49 SUBUNITS IN THREE TURNS. A SINGLE STRAND OF RNA FOLLOWS THE BASIC HELIX BETWEEN THE PROTEIN SUBUNITS AT A DISTANCE OF 40 ANGSTROMS. THERE ARE THREE NUCLEOTIDES BOUND TO EACH PROTEIN SUBUNIT. |
- Components
Components
| #1: RNA chain | Mass: 958.660 Da / Num. of mol.: 1 / Source method: obtained synthetically | ||
|---|---|---|---|
| #2: Protein | Mass: 17505.426 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   Tobacco mosaic virus / Strain: vulgare / References: UniProt: P03570, UniProt: P69687*PLUS Tobacco mosaic virus / Strain: vulgare / References: UniProt: P03570, UniProt: P69687*PLUS | ||
| #3: Chemical | ChemComp-CA / | ||
| #4: Water | ChemComp-HOH / | ||
| Nonpolymer details | WATER MOLECULE 45 OCCUPIES A SITE BELIEVED TO BIND CALCIUM AT HIGHER CALCIUM CONCENTRAT| Sequence details | GENERATION OF DIFFERENT SUBUNITS IS MOST EASILY DONE IN POLAR COORDINATES. ASSUME THAT THE ...GENERATION | |
-Experimental details
-Experiment
| Experiment | Method:  FIBER DIFFRACTION FIBER DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal grow | *PLUS Method: unknown |
|---|
- Processing
Processing
| Software | Name: PROLSQ / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Rfactor obs: 0.096 / Highest resolution: 2.9 Å Details: THE STRUCTURE WAS DETERMINED BY FIBER DIFFRACTION USING MULTI-DIMENSIONAL ISOMORPHOUS REPLACEMENT WITH LAYER-LINE SPLITTING, SOLVENT FLATTENING REFINEMENT AND RESTRAINED LEAST SQUARES ...Details: THE STRUCTURE WAS DETERMINED BY FIBER DIFFRACTION USING MULTI-DIMENSIONAL ISOMORPHOUS REPLACEMENT WITH LAYER-LINE SPLITTING, SOLVENT FLATTENING REFINEMENT AND RESTRAINED LEAST SQUARES COORDINATE REFINEMENT. THE STRUCTURE INCLUDES 154 OF THE 158 AMINO ACIDS AND THREE RNA NUCLEOTIDES MODELLED AS GAA BUT REPRESENTING THE ENTIRE NUCLEIC ACID CONTENT. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Highest resolution: 2.9 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
|
 Movie
Movie Controller
Controller







 PDBj
PDBj



