+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1rmv | ||||||
|---|---|---|---|---|---|---|---|
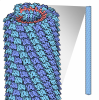
| Title | RIBGRASS MOSAIC VIRUS, FIBER DIFFRACTION | ||||||
 Components Components |
| ||||||
 Keywords Keywords | Virus/RNA / RIBGRASS MOSAIC VIRUS / TOBAMOVIRUS / RMV CLUSTER / COAT PROTEIN (VIRAL) / COMPLEX (COAT PROTEIN-RNA) / Helical virus / Virus-RNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Ribgrass mosaic virus Ribgrass mosaic virus | ||||||
| Method |  FIBER DIFFRACTION / FIBER DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.9 Å MOLECULAR REPLACEMENT / Resolution: 2.9 Å | ||||||
 Authors Authors | Wang, H. / Stubbs, G. | ||||||
 Citation Citation |  Journal: Acta Crystallogr.,Sect.A / Year: 1993 Journal: Acta Crystallogr.,Sect.A / Year: 1993Title: Molecular dynamics in refinement against fiber diffraction data. Authors: Wang, H. / Stubbs, G. #1:  Journal: J.Mol.Biol. / Year: 1993 Journal: J.Mol.Biol. / Year: 1993Title: Preliminary X-Ray Diffraction Studies of Ribgrass Mosaic Virus Authors: Wang, H. / Planchart, A. / Allen, D. / Pattanayek, R. / Stubbs, G. #2:  Journal: Acta Crystallogr.,Sect.A / Year: 1985 Journal: Acta Crystallogr.,Sect.A / Year: 1985Title: Solving the Phase Problem in Fiber Diffraction. Application to Tobacco Mosaic Virus at 3.6 Angstroms Resolution Authors: Namba, K. / Stubbs, G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1rmv.cif.gz 1rmv.cif.gz | 46.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1rmv.ent.gz pdb1rmv.ent.gz | 32.6 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1rmv.json.gz 1rmv.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rm/1rmv https://data.pdbj.org/pub/pdb/validation_reports/rm/1rmv ftp://data.pdbj.org/pub/pdb/validation_reports/rm/1rmv ftp://data.pdbj.org/pub/pdb/validation_reports/rm/1rmv | HTTPS FTP |
|---|
-Related structure data
| Related structure data |  2tmvS S: Starting model for refinement |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | x 49
| ||||||||
| 2 |
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
| ||||||||
| Symmetry | Helical symmetry: (Circular symmetry: 1 / Dyad axis: no / N subunits divisor: 49 / Num. of operations: 49 / Rise per n subunits: 69.7 Å / Rotation per n subunits: 1080 °) | ||||||||
| Details | THE BASIC REPEAT UNIT OF THE VIRAL HELIX IS 69.7 ANGSTROMS IN LENGTH WITH 49 SUBUNITS IN 3 HELICAL TURNS. |
- Components
Components
| #1: RNA chain | Mass: 958.660 Da / Num. of mol.: 1 / Fragment: GAA / Source method: isolated from a natural source / Source: (natural)  Ribgrass mosaic virus / Genus: Tobamovirus Ribgrass mosaic virus / Genus: Tobamovirus |
|---|---|
| #2: Protein | Mass: 17552.461 Da / Num. of mol.: 1 / Source method: isolated from a natural source / Source: (natural)  Ribgrass mosaic virus / Genus: Tobamovirus / References: UniProt: P03580 Ribgrass mosaic virus / Genus: Tobamovirus / References: UniProt: P03580 |
| Has protein modification | Y |
-Experimental details
-Experiment
| Experiment | Method:  FIBER DIFFRACTION FIBER DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal grow | pH: 7.2 / Details: pH 7.2 |
|---|---|
| Crystal grow | *PLUS |
-Data collection
| Diffraction | Mean temperature: 298 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ELLIOT GX-6 / Wavelength: 1.5418 ROTATING ANODE / Type: ELLIOT GX-6 / Wavelength: 1.5418 |
| Detector | Type: KODAK FILM / Detector: FILM / Date: Oct 1, 1990 / Details: MIRRORS |
| Radiation | Monochromatic (M) / Laue (L): M |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.8→50 Å / Num. obs: 6045 / % possible obs: 91.2 % / Redundancy: 4 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: PDB ENTRY 2TMV Resolution: 2.9→10 Å / σ(F): 0
| ||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.9→10 Å
| ||||||||||||||||||||
| Software | *PLUS Name: FXPLOR / Classification: refinement | ||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.095 | ||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller










 PDBj
PDBj
