[English] 日本語
 Yorodumi
Yorodumi- PDB-1vtm: STRUCTURE OF THE U2 STRAIN OF TOBACCO MOSAIC VIRUS REFINED AT 3.5... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1vtm | ||||||
|---|---|---|---|---|---|---|---|
| Title | STRUCTURE OF THE U2 STRAIN OF TOBACCO MOSAIC VIRUS REFINED AT 3.5 ANGSTROMS RESOLUTION USING X-RAY FIBER DIFFRACTION | ||||||
 Components Components |
| ||||||
 Keywords Keywords | Virus/RNA / VIRUS / Helical virus / Virus-RNA COMPLEX | ||||||
| Function / homology |  Function and homology information Function and homology information | ||||||
| Biological species |  Tobacco mild green mosaic virus Tobacco mild green mosaic virus | ||||||
| Method |  FIBER DIFFRACTION / Resolution: 3.5 Å FIBER DIFFRACTION / Resolution: 3.5 Å | ||||||
 Authors Authors | Stubbs, G. / Pattanayek, R. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1992 Journal: J.Mol.Biol. / Year: 1992Title: Structure of the U2 strain of tobacco mosaic virus refined at 3.5 A resolution using X-ray fiber diffraction. Authors: Pattanayek, R. / Stubbs, G. #1:  Journal: Acta Crystallogr.,Sect.A / Year: 1993 Journal: Acta Crystallogr.,Sect.A / Year: 1993Title: Molecular Dynamics in Refinement Against Fiber Diffraction Data Authors: Wang, H. / Stubbs, G. #2:  Journal: Biophys.J. / Year: 1986 Journal: Biophys.J. / Year: 1986Title: Application of Restrained Least-Squares Refinement to Fiber Diffraction from Macromolecular Assemblies Authors: Stubbs, G. / Namba, K. / Makowski, L. #3:  Journal: Acta Crystallogr.,Sect.A / Year: 1985 Journal: Acta Crystallogr.,Sect.A / Year: 1985Title: Solving the Phase Problem in Fiber Diffraction. Application to Tobacco Mosaic Virus at 3.6 Angstroms Resolution Authors: Namba, K. / Stubbs, G. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1vtm.cif.gz 1vtm.cif.gz | 46.9 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1vtm.ent.gz pdb1vtm.ent.gz | 33.2 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1vtm.json.gz 1vtm.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/vt/1vtm https://data.pdbj.org/pub/pdb/validation_reports/vt/1vtm ftp://data.pdbj.org/pub/pdb/validation_reports/vt/1vtm ftp://data.pdbj.org/pub/pdb/validation_reports/vt/1vtm | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | x 49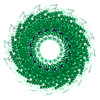
| ||||||||
| 2 |
| ||||||||
| 3 | 
| ||||||||
| Unit cell |
| ||||||||
| Atom site foot note | 1: GLN P 99 - PRO P 100 OMEGA = 251.92 PEPTIDE BOND DEVIATES SIGNIFICANTLY FROM TRANS CONFORMATION | ||||||||
| Symmetry | Helical symmetry: (Circular symmetry: 1 / Dyad axis: no / N subunits divisor: 49 / Num. of operations: 49 / Rise per n subunits: 69 Å / Rotation per n subunits: 1080 °) | ||||||||
| Details | THE TRANSFORMATION PRESENTED ON *MTRIX* RECORDS BELOW REPRESENTS HELICAL SYMMETRY WITH THE HELIX AXIS ON THE Z AXIS. THERE ARE 49 SUBUNITS IN 3 TURNS OF THE HELIX. THE FULL (49 SUBUNIT) HELICAL REPEAT IS 69 ANGSTROMS. |
- Components
Components
| #1: RNA chain | Mass: 958.660 Da / Num. of mol.: 1 / Source method: obtained synthetically Source: (gene. exp.)  Tobacco mild green mosaic virus (TMGMV) Tobacco mild green mosaic virus (TMGMV)Strain: U2 / Gene: CP |
|---|---|
| #2: Protein | Mass: 17467.326 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source References: UniProt: P03579 |
| #3: Water | ChemComp-HOH / |
| Sequence details | CONCERNING THE APPARENT SEQUENCE DISCREPANCY: THE AUTHORS USED THE SEQUENCE REPORTED BY ALTSCHUH ET ...CONCERNING |
-Experimental details
-Experiment
| Experiment | Method:  FIBER DIFFRACTION FIBER DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal grow | *PLUS Method: unknown |
|---|
- Processing
Processing
| Software |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 3.5→10 Å / Rfactor obs: 0.096 Details: THIS STRUCTURE WAS DETERMINED FROM FIBER DIFFRACTION DATA BY MOLECULAR REPLACEMENT FROM TMV, PROTEIN DATA BANK ENTRY 2TMV. U2 HAS 72 PER CENT SEQUENCE HOMOLOGY WITH TMV. THIS STRUCTURE ...Details: THIS STRUCTURE WAS DETERMINED FROM FIBER DIFFRACTION DATA BY MOLECULAR REPLACEMENT FROM TMV, PROTEIN DATA BANK ENTRY 2TMV. U2 HAS 72 PER CENT SEQUENCE HOMOLOGY WITH TMV. THIS STRUCTURE INCLUDES ALL 158 AMINO ACIDS AND 3 RNA NUCLEOTIDES, MODELED AS GAA BUT REPRESENTING THE ENTIRE GENOME. THERE IS ONE SMALL SHEET IN THIS STRUCTURE BUT IT IS TOO IRREGULAR TO INCLUDE. | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 3.5→10 Å
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.096 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | *PLUS Highest resolution: 3.53 Å / Lowest resolution: 3.77 Å / Total num. of bins used: 7 / Rfactor obs: 0.171 |
 Movie
Movie Controller
Controller









 PDBj
PDBj

