+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: PDB / ID: 2jua | ||||||
|---|---|---|---|---|---|---|---|
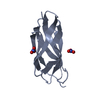
| タイトル | Assignment, structure, and dynamics of de novo designed protein S836 | ||||||
 要素 要素 | de novo protein S836 | ||||||
 キーワード キーワード | DE NOVO PROTEIN | ||||||
| 機能・相同性 | Designed four-helix bundle protein / hypothetical protein mp506/mpn330, domain 1 / Up-down Bundle / Mainly Alpha 機能・相同性情報 機能・相同性情報 | ||||||
| 生物種 | unidentified (未定義) | ||||||
| 手法 | 溶液NMR / simulated annealing | ||||||
 データ登録者 データ登録者 | Go, A. / Kim, S. / Baum, J.S. / Hecht, M.H. | ||||||
 引用 引用 |  ジャーナル: Protein Sci. / 年: 2008 ジャーナル: Protein Sci. / 年: 2008タイトル: Structure and dynamics of de novo proteins from a designed superfamily of 4-helix bundles. 著者: Go, A. / Kim, S. / Baum, J. / Hecht, M.H. #1:  ジャーナル: To be Published ジャーナル: To be Publishedタイトル: NMR assignment of S836: a de novo Protein from a Designed Protein Superfamily 著者: Go, A. / Kim, S. / Hecht, M.H. / Baum, J.S. | ||||||
| 履歴 |
| ||||||
| Remark 999 | Sequence This is a de novo Protein design sequence. |
- 構造の表示
構造の表示
| 構造ビューア | 分子:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- ダウンロードとリンク
ダウンロードとリンク
- ダウンロード
ダウンロード
| PDBx/mmCIF形式 |  2jua.cif.gz 2jua.cif.gz | 639.7 KB | 表示 |  PDBx/mmCIF形式 PDBx/mmCIF形式 |
|---|---|---|---|---|
| PDB形式 |  pdb2jua.ent.gz pdb2jua.ent.gz | 533.2 KB | 表示 |  PDB形式 PDB形式 |
| PDBx/mmJSON形式 |  2jua.json.gz 2jua.json.gz | ツリー表示 |  PDBx/mmJSON形式 PDBx/mmJSON形式 | |
| その他 |  その他のダウンロード その他のダウンロード |
-検証レポート
| 文書・要旨 |  2jua_validation.pdf.gz 2jua_validation.pdf.gz | 339.2 KB | 表示 |  wwPDB検証レポート wwPDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  2jua_full_validation.pdf.gz 2jua_full_validation.pdf.gz | 532.4 KB | 表示 | |
| XML形式データ |  2jua_validation.xml.gz 2jua_validation.xml.gz | 64.4 KB | 表示 | |
| CIF形式データ |  2jua_validation.cif.gz 2jua_validation.cif.gz | 85.1 KB | 表示 | |
| アーカイブディレクトリ |  https://data.pdbj.org/pub/pdb/validation_reports/ju/2jua https://data.pdbj.org/pub/pdb/validation_reports/ju/2jua ftp://data.pdbj.org/pub/pdb/validation_reports/ju/2jua ftp://data.pdbj.org/pub/pdb/validation_reports/ju/2jua | HTTPS FTP |
-関連構造データ
| 関連構造データ | |
|---|---|
| 類似構造データ | |
| その他のデータベース |
|
- リンク
リンク
- 集合体
集合体
| 登録構造単位 | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR アンサンブル |
|
- 要素
要素
| #1: タンパク質 | 分子量: 11946.391 Da / 分子数: 1 / 由来タイプ: 組換発現 詳細: Ambiguous conformational states: Y2, Q24, and N84 register possible alternate alpha carbon chemical shifts 由来: (組換発現) unidentified (未定義) / プラスミド: pet3A / 発現宿主:  |
|---|
-実験情報
-実験
| 実験 | 手法: 溶液NMR | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| NMR実験 |
|
- 試料調製
試料調製
| 詳細 |
| ||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 試料 |
| ||||||||||||||||||||||||||||
| 試料状態 | pH: 4 / 圧: ambient / 温度: 298 K |
-NMR測定
| NMRスペクトロメーター |
|
|---|
- 解析
解析
| NMR software |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 精密化 | 手法: simulated annealing / ソフトェア番号: 1 | ||||||||||||||||||
| 代表構造 | 選択基準: lowest energy | ||||||||||||||||||
| NMRアンサンブル | コンフォーマー選択の基準: structures with the lowest energy 計算したコンフォーマーの数: 100 / 登録したコンフォーマーの数: 20 |
 ムービー
ムービー コントローラー
コントローラー













 PDBj
PDBj
 HSQC
HSQC