[English] 日本語
 Yorodumi
Yorodumi- PDB-280d: THE STRUCTURE OF AN RNA DODECAMER SHOWS HOW TANDEM U-U BASE PAIRS... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 280d | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
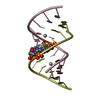
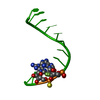
| Title | THE STRUCTURE OF AN RNA DODECAMER SHOWS HOW TANDEM U-U BASE PAIRS INCREASE THE RANGE OF STABLE RNA STRUCTURES AND THE DIVERSITY OF RECOGNITION SITES | ||||||||||||||||||
 Components Components | RNA (5'-R(* Keywords KeywordsRNA / UNUSUAL RNA / DOUBLE HELIX / INTERNAL LOOP / MISMATCHED | Function / homology | RNA / RNA (> 10) |  Function and homology information Function and homology informationMethod |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  MOLECULAR REPLACEMENT / Resolution: 2.4 Å MOLECULAR REPLACEMENT / Resolution: 2.4 Å  Authors AuthorsLietzke, S.E. / Barnes, C.L. / Kundrot, C.E. |  Citation Citation Journal: Structure / Year: 1996 Journal: Structure / Year: 1996Title: The structure of an RNA dodecamer shows how tandem U-U base pairs increase the range of stable RNA structures and the diversity of recognition sites. Authors: Lietzke, S.E. / Barnes, C.L. / Berglund, J.A. / Kundrot, C.E. #1:  Journal: Acta Crystallogr.,Sect.D / Year: 1996 Journal: Acta Crystallogr.,Sect.D / Year: 1996Title: Data reduction from twinned RNA crystals Authors: Lietzke, S.E. / Carperos, V.E. / Kundrot, C.E. #2:  Journal: Proc.Natl.Acad.Sci.USA / Year: 1993 Journal: Proc.Natl.Acad.Sci.USA / Year: 1993Title: Crystallization of Ribozymes and Small RNA Motifs by a Sparse Matrix Approach Authors: Doudna, J.A. / Grosshans, C. / Gooding, A. / Kundrot, C.E. History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  280d.cif.gz 280d.cif.gz | 39.2 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb280d.ent.gz pdb280d.ent.gz | 27 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  280d.json.gz 280d.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/80/280d https://data.pdbj.org/pub/pdb/validation_reports/80/280d ftp://data.pdbj.org/pub/pdb/validation_reports/80/280d ftp://data.pdbj.org/pub/pdb/validation_reports/80/280d | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
|
- Components
Components
| #1: RNA chain | Mass: 3820.296 Da / Num. of mol.: 4 Source method: isolated from a genetically manipulated source Description: THE DODECAMER WAS PREPARED BY TRANSCRIPTION WITH T7 RNA POLYMERASE FROM A SINGLE STRANDED DNA TEMPLATE. Plasmid: PAR1219 / Species (production host): Escherichia coli / Production host:  #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 2 X-RAY DIFFRACTION / Number of used crystals: 2 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.44 Å3/Da / Density % sol: 49.57 % Description: THE CRYSTALS WERE TWINNED; DATA FROM EACH LATTICE WAS INDIVIDUALLY PROCESSED WITH XDS AND A SET OF PROGRAMS DESCRIBED IN REFERENCE 1. DATA FROM THE TWO LATTICES WERE THEN MERGED WITH XSCALE. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | Method: vapor diffusion / pH: 7 / Details: pH 7.00, VAPOR DIFFUSION / Temp details: ROOM TEMPERATURE | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Crystal grow | *PLUS pH: 7 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 100 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 ROTATING ANODE / Type: RIGAKU RU200 / Wavelength: 1.5418 |
| Detector | Type: SIEMENS-NICOLET X100 / Detector: AREA DETECTOR / Date: Mar 22, 1995 |
| Radiation | Monochromator: GRAPHITE / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.4→15 Å / Num. obs: 5429 / % possible obs: 95.4 % / Redundancy: 4.9 % / Rmerge(I) obs: 0.076 / Net I/σ(I): 44.6 |
| Reflection shell | Resolution: 2.4→2.6 Å / Redundancy: 2.4 % / Rmerge(I) obs: 0.182 / Mean I/σ(I) obs: 8.05 / % possible all: 95.5 |
| Reflection | *PLUS Highest resolution: 2.4 Å / Lowest resolution: 15 Å / % possible obs: 95.4 % / Redundancy: 4.9 % / Num. measured all: 26632 |
| Reflection shell | *PLUS Highest resolution: 2.4 Å / Lowest resolution: 2.6 Å / Redundancy: 2.4 % / Mean I/σ(I) obs: 8.05 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT MOLECULAR REPLACEMENTStarting model: HELIX 1 OF THE RNA DODECAMER CONTAINING THE E. COLI SHINE DALGARNO SEQUENCE (NDB ID ARL062) WITH THE SEQUENCE MUTATED TO R(GGCGCUUGCGUC). Resolution: 2.4→8 Å / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine Biso |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.4→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| LS refinement shell | Resolution: 2.4→2.51 Å /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Xplor file |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Highest resolution: 2.4 Å / Lowest resolution: 8 Å / σ(F): 2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller







 PDBj
PDBj






























