[English] 日本語
 Yorodumi
Yorodumi- PDB-1ytc: THERMODYNAMIC CYCLES AS PROBES OF STRUCTURE-FUNCTION RELATIONSHIP... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ytc | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
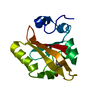
| Title | THERMODYNAMIC CYCLES AS PROBES OF STRUCTURE-FUNCTION RELATIONSHIPS IN UNFOLDED PROTEINS | |||||||||
 Components Components | YEAST ISO-2 CYTOCHROME C | |||||||||
 Keywords Keywords | ELECTRON TRANSPORT / HEME PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationRelease of apoptotic factors from the mitochondria / Pyroptosis / Detoxification of Reactive Oxygen Species / Respiratory electron transport / mitochondrial electron transport, cytochrome c to oxygen / mitochondrial electron transport, ubiquinol to cytochrome c / mitochondrial intermembrane space / electron transfer activity / heme binding / mitochondrion / metal ion binding Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method |  X-RAY DIFFRACTION / Resolution: 1.8 Å X-RAY DIFFRACTION / Resolution: 1.8 Å | |||||||||
 Authors Authors | Luo, Y. / Brayer, G.D. | |||||||||
 Citation Citation |  Journal: Biochemistry / Year: 1996 Journal: Biochemistry / Year: 1996Title: Thermodynamic cycles as probes of structure in unfolded proteins. Authors: McGee, W.A. / Rosell, F.I. / Liggins, J.R. / Rodriguez-Ghidarpour, S. / Luo, Y. / Chen, J. / Brayer, G.D. / Mauk, A.G. / Nall, B.T. #1:  Journal: Protein Sci. / Year: 1993 Journal: Protein Sci. / Year: 1993Title: The Structure and Function of Omega Loop a Replacements in Cytochrome C Authors: Murphy, M.E.P. / Fetrow, J.S. / Burton, R.E. / Brayer, G.D. #2:  Journal: J.Mol.Biol. / Year: 1992 Journal: J.Mol.Biol. / Year: 1992Title: Structure Determination and Analysis of Yeast Iso-2-Cytochrome C and a Composite Mutant Protein Authors: Murphy, M.E.P. / Nall, B.T. / Brayer, G.D. | |||||||||
| History |
| |||||||||
| Remark 700 | SHEET THE PDB HAS GENERATED THE HELIX AND SHEET RECORDS FROM THE COORDINATES USING A PROGRAM BASED ...SHEET THE PDB HAS GENERATED THE HELIX AND SHEET RECORDS FROM THE COORDINATES USING A PROGRAM BASED ON DSSP OF W. KABSCH AND C. SANDER. |
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ytc.cif.gz 1ytc.cif.gz | 35.6 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ytc.ent.gz pdb1ytc.ent.gz | 26.7 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ytc.json.gz 1ytc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/yt/1ytc https://data.pdbj.org/pub/pdb/validation_reports/yt/1ytc ftp://data.pdbj.org/pub/pdb/validation_reports/yt/1ytc ftp://data.pdbj.org/pub/pdb/validation_reports/yt/1ytc | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 12465.292 Da / Num. of mol.: 1 / Mutation: N52I Source method: isolated from a genetically manipulated source Details: REDUCED STATE OF HEME Source: (gene. exp.)  Production host:  |
|---|---|
| #2: Chemical | ChemComp-SO4 / |
| #3: Chemical | ChemComp-HEC / |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 1.88 Å3/Da / Density % sol: 34.5 % | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS pH: 7 / Method: unknown | ||||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction source | Wavelength: 1.5418 Å |
|---|---|
| Detector | Type: RIGAKU / Detector: IMAGE PLATE / Date: Mar 24, 1992 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 1.8→69.5 Å / Num. obs: 9141 / % possible obs: 95.5 % / Observed criterion σ(I): 2 / Redundancy: 6 % / Rmerge(I) obs: 0.06 |
| Reflection | *PLUS Lowest resolution: 9999 Å / Num. measured all: 59573 / Rmerge(I) obs: 0.06 |
| Reflection shell | *PLUS Highest resolution: 1.8 Å / Lowest resolution: 2 Å / % possible obs: 93.4 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 1.8→8 Å / σ(F): 2
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 24 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.16 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 1.8→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller











 PDBj
PDBj












