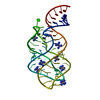
Entry Database : PDB / ID : 1y27Title G-riboswitch-guanine complex Bacillus subtilis xpt Keywords / Function / homology / / Biological species Bacillus subtilis (bacteria)Method / / / Resolution : 2.4 Å Authors Serganov, A. / Yuan, Y.R. / Patel, D.J. Journal : Chem.Biol. / Year : 2004Title : Structural Basis for Discriminative Regulation of Gene Expression by Adenine- and Guanine-Sensing mRNAsAuthors : Serganov, A. / Yuan, Y.R. / Pikovskaya, O. / Polonskaia, A. / Malinina, L. / Phan, A.T. / Hobartner, C. / Micura, R. / Breaker, R.R. / Patel, D.J. History Deposition Nov 20, 2004 Deposition site / Processing site Revision 1.0 Dec 28, 2004 Provider / Type Revision 1.1 Apr 30, 2008 Group Revision 1.2 Jul 13, 2011 Group / Version format complianceRevision 1.3 Oct 11, 2017 Group / Category / Item / _software.nameRevision 2.0 Feb 27, 2019 Group Advisory / Atomic model ... Advisory / Atomic model / Data collection / Database references / Derived calculations / Non-polymer description / Polymer sequence / Refinement description / Source and taxonomy / Structure summary Category atom_site / chem_comp ... atom_site / chem_comp / entity / entity_name_com / entity_poly / entity_poly_seq / entity_src_gen / ndb_struct_na_base_pair / ndb_struct_na_base_pair_step / pdbx_entity_nonpoly / pdbx_nonpoly_scheme / pdbx_poly_seq_scheme / pdbx_refine_tls_group / pdbx_struct_assembly / pdbx_struct_assembly_gen / pdbx_unobs_or_zero_occ_atoms / pdbx_validate_close_contact / pdbx_validate_planes / pdbx_validate_rmsd_angle / pdbx_validate_rmsd_bond / refine / refine_hist / refine_ls_restr / refine_ls_shell / reflns / reflns_shell / struct_asym / struct_conn / struct_ref / struct_site / struct_site_gen Item _atom_site.B_iso_or_equiv / _atom_site.Cartn_x ... _atom_site.B_iso_or_equiv / _atom_site.Cartn_x / _atom_site.Cartn_y / _atom_site.Cartn_z / _atom_site.auth_atom_id / _atom_site.auth_comp_id / _atom_site.auth_seq_id / _atom_site.group_PDB / _atom_site.label_asym_id / _atom_site.label_atom_id / _atom_site.label_comp_id / _atom_site.label_entity_id / _atom_site.label_seq_id / _atom_site.type_symbol / _chem_comp.formula / _chem_comp.formula_weight / _chem_comp.id / _chem_comp.mon_nstd_flag / _chem_comp.name / _chem_comp.type / _entity_poly.pdbx_seq_one_letter_code / _entity_poly_seq.mon_id / _entity_src_gen.pdbx_beg_seq_num / _entity_src_gen.pdbx_end_seq_num / _entity_src_gen.pdbx_seq_type / _ndb_struct_na_base_pair.buckle / _ndb_struct_na_base_pair.hbond_type_12 / _ndb_struct_na_base_pair.hbond_type_28 / _ndb_struct_na_base_pair.i_auth_seq_id / _ndb_struct_na_base_pair.i_label_comp_id / _ndb_struct_na_base_pair.i_label_seq_id / _ndb_struct_na_base_pair.j_auth_seq_id / _ndb_struct_na_base_pair.j_label_comp_id / _ndb_struct_na_base_pair.j_label_seq_id / _ndb_struct_na_base_pair.opening / _ndb_struct_na_base_pair.pair_name / _ndb_struct_na_base_pair.propeller / _ndb_struct_na_base_pair.shear / _ndb_struct_na_base_pair.stagger / _ndb_struct_na_base_pair.stretch / _pdbx_poly_seq_scheme.auth_mon_id / _pdbx_poly_seq_scheme.mon_id / _pdbx_poly_seq_scheme.pdb_mon_id / _pdbx_refine_tls_group.beg_label_asym_id / _pdbx_refine_tls_group.beg_label_seq_id / _pdbx_refine_tls_group.end_label_asym_id / _pdbx_refine_tls_group.end_label_seq_id / _pdbx_struct_assembly.details / _pdbx_struct_assembly.method_details / _pdbx_struct_assembly_gen.asym_id_list / _refine.B_iso_mean / _refine.details / _refine.ls_R_factor_R_free / _refine.ls_R_factor_R_work / _refine.ls_R_factor_obs / _refine.ls_percent_reflns_obs / _refine.pdbx_ls_sigma_F / _refine_hist.pdbx_number_atoms_ligand / _refine_hist.pdbx_number_atoms_nucleic_acid / _refine_ls_shell.d_res_low / _reflns.d_resolution_low / _reflns.number_all / _reflns.observed_criterion_sigma_F / _reflns.percent_possible_obs / _reflns_shell.number_unique_all / _reflns_shell.percent_possible_all / _struct_ref.pdbx_align_begin Revision 2.1 Aug 23, 2023 Group / Database references / Refinement descriptionCategory chem_comp_atom / chem_comp_bond ... chem_comp_atom / chem_comp_bond / database_2 / pdbx_initial_refinement_model Item / _database_2.pdbx_database_accession
Show all Show less Remark 999 SEQUENCE THE G-RIBOSWITCH SEQUENCE WAS DERIVED FROM THE SEQUENCE LOCATED UPSTREAM OF THE XPT GENE ... SEQUENCE THE G-RIBOSWITCH SEQUENCE WAS DERIVED FROM THE SEQUENCE LOCATED UPSTREAM OF THE XPT GENE (XPT-PBUX OPERON) ENCODING FOR THE XANTHINE PHOPHORIBOSYLTRANSFERASE FROM BACILLUS SUBTILIS. THE STRUCTURE IN THIS ENTRY IS A COMPLEX BETWEEN GUANINE AND A DOMAIN (APTAMER DOMAIN) OF THE RIBOSWITCH, WHICH SHUTS DOWN TRANSCRIPTION OF THE XPT GENE IN RESPONSE TO THE PRESENCE OF PURINES IN THE CELL. THIS SEQUENCE CAN BE FOUND NATURALLY AND THE DATABASE MATCH IS FOUND IN BACILLUS SUBTILIS GENEBANK ENTRY 633168 X83878 (RESIDUES 185-252).
 Open data
Open data Basic information
Basic information Components
Components Keywords
Keywords Function and homology information
Function and homology information
 X-RAY DIFFRACTION /
X-RAY DIFFRACTION /  SYNCHROTRON /
SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2.4 Å
MOLECULAR REPLACEMENT / Resolution: 2.4 Å  Authors
Authors Citation
Citation Journal: Chem.Biol. / Year: 2004
Journal: Chem.Biol. / Year: 2004 Structure visualization
Structure visualization Molmil
Molmil Jmol/JSmol
Jmol/JSmol Downloads & links
Downloads & links Download
Download 1y27.cif.gz
1y27.cif.gz PDBx/mmCIF format
PDBx/mmCIF format pdb1y27.ent.gz
pdb1y27.ent.gz PDB format
PDB format 1y27.json.gz
1y27.json.gz PDBx/mmJSON format
PDBx/mmJSON format Other downloads
Other downloads https://data.pdbj.org/pub/pdb/validation_reports/y2/1y27
https://data.pdbj.org/pub/pdb/validation_reports/y2/1y27 ftp://data.pdbj.org/pub/pdb/validation_reports/y2/1y27
ftp://data.pdbj.org/pub/pdb/validation_reports/y2/1y27
 Links
Links Assembly
Assembly
 Components
Components

 X-RAY DIFFRACTION / Number of used crystals: 1
X-RAY DIFFRACTION / Number of used crystals: 1  Sample preparation
Sample preparation SYNCHROTRON / Site:
SYNCHROTRON / Site:  NSLS
NSLS  / Beamline: X25 / Wavelength: 1.14 Å
/ Beamline: X25 / Wavelength: 1.14 Å Processing
Processing MOLECULAR REPLACEMENT
MOLECULAR REPLACEMENT Movie
Movie Controller
Controller










 PDBj
PDBj