+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1rcp | ||||||
|---|---|---|---|---|---|---|---|
| Title | CYTOCHROME C' | ||||||
 Components Components | CYTOCHROME C' | ||||||
 Keywords Keywords | ELECTRON TRANSPORT / CYTOCHROME / HEME | ||||||
| Function / homology |  Function and homology information Function and homology informationelectron transport chain / periplasmic space / electron transfer activity / iron ion binding / heme binding Similarity search - Function | ||||||
| Biological species |  Rhodobacter capsulatus (bacteria) Rhodobacter capsulatus (bacteria) | ||||||
| Method |  X-RAY DIFFRACTION / X-RAY DIFFRACTION /  SYNCHROTRON / SYNCHROTRON /  MOLECULAR REPLACEMENT / Resolution: 2 Å MOLECULAR REPLACEMENT / Resolution: 2 Å | ||||||
 Authors Authors | Tahirov, T.H. / Misaki, S. / Meyer, T.E. / Cusanovich, M.A. / Higuchi, Y. / Yasuoka, N. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: High-resolution crystal structures of two polymorphs of cytochrome c' from the purple phototrophic bacterium rhodobacter capsulatus. Authors: Tahirov, T.H. / Misaki, S. / Meyer, T.E. / Cusanovich, M.A. / Higuchi, Y. / Yasuoka, N. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1rcp.cif.gz 1rcp.cif.gz | 75.5 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1rcp.ent.gz pdb1rcp.ent.gz | 58 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1rcp.json.gz 1rcp.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/rc/1rcp https://data.pdbj.org/pub/pdb/validation_reports/rc/1rcp ftp://data.pdbj.org/pub/pdb/validation_reports/rc/1rcp ftp://data.pdbj.org/pub/pdb/validation_reports/rc/1rcp | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 | 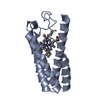
| ||||||||
| 2 | 
| ||||||||
| Unit cell |
| ||||||||
| Noncrystallographic symmetry (NCS) | NCS oper: (Code: given Matrix: (-0.99999, -0.00113, -0.00433), Vector: |
- Components
Components
| #1: Protein | Mass: 13154.733 Da / Num. of mol.: 2 / Source method: isolated from a natural source Details: STRAIN M110 WAS DERIVED FROM THE WILD TYPE GENETIC STRAIN ST. LOUIS. THE CYTOCHROME C' DIFFERS FROM STRAIN SP7 IN 12 RESIDUES Source: (natural)  Rhodobacter capsulatus (bacteria) / Strain: MT110 / References: UniProt: P00147 Rhodobacter capsulatus (bacteria) / Strain: MT110 / References: UniProt: P00147#2: Chemical | #3: Water | ChemComp-HOH / | Has protein modification | Y | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.78 Å3/Da / Density % sol: 53 % | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 6.5 / Details: pH 6.5 | |||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion, sitting drop | |||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 283 K |
|---|---|
| Diffraction source | Source:  SYNCHROTRON / Site: SYNCHROTRON / Site:  Photon Factory Photon Factory  / Beamline: BL-6A / Wavelength: 1 / Beamline: BL-6A / Wavelength: 1 |
| Detector | Type: FUJI / Detector: IMAGE PLATE / Date: Dec 22, 1994 |
| Radiation | Monochromator: Y / Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1 Å / Relative weight: 1 |
| Reflection | Resolution: 2→30 Å / Num. obs: 15826 / % possible obs: 79.3 % / Observed criterion σ(I): -3 / Redundancy: 3.89 % / Rmerge(I) obs: 0.057 / Net I/σ(I): 20.09 |
| Reflection | *PLUS Num. measured all: 61516 |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure:  MOLECULAR REPLACEMENT / Resolution: 2→6 Å / σ(F): 2 / MOLECULAR REPLACEMENT / Resolution: 2→6 Å / σ(F): 2 /
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | Biso mean: 25.01 Å2 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine analyze | Luzzati coordinate error obs: 0.18 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2→6 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name:  X-PLOR / Classification: refinement X-PLOR / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller














 PDBj
PDBj








