[English] 日本語
 Yorodumi
Yorodumi- PDB-1qlc: Solution structure of the second PDZ domain of Postsynaptic Density-95 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1qlc | ||||||
|---|---|---|---|---|---|---|---|
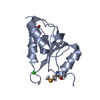
| Title | Solution structure of the second PDZ domain of Postsynaptic Density-95 | ||||||
 Components Components | POSTSYNAPTIC DENSITY PROTEIN 95 | ||||||
 Keywords Keywords | PEPTIDE RECOGNITION / PDZ DOMAIN / NEURONAL NITRIC OXIDE SYNTHASE / NMDA RECEPTOR BINDING | ||||||
| Function / homology |  Function and homology information Function and homology informationRHO GTPases activate CIT / neuronal ion channel clustering / positive regulation of AMPA glutamate receptor clustering / P2Y1 nucleotide receptor binding / Neurexins and neuroligins / beta-1 adrenergic receptor binding / neuroligin family protein binding / regulation of grooming behavior / structural constituent of postsynaptic density / synaptic vesicle maturation ...RHO GTPases activate CIT / neuronal ion channel clustering / positive regulation of AMPA glutamate receptor clustering / P2Y1 nucleotide receptor binding / Neurexins and neuroligins / beta-1 adrenergic receptor binding / neuroligin family protein binding / regulation of grooming behavior / structural constituent of postsynaptic density / synaptic vesicle maturation / positive regulation of neuron projection arborization / receptor localization to synapse / cerebellar mossy fiber / protein localization to synapse / LGI-ADAM interactions / neuron spine / dendritic spine morphogenesis / Trafficking of AMPA receptors / proximal dendrite / negative regulation of receptor internalization / juxtaparanode region of axon / Activation of Ca-permeable Kainate Receptor / vocalization behavior / dendritic branch / acetylcholine receptor binding / positive regulation of dendrite morphogenesis / frizzled binding / cellular response to potassium ion / regulation of NMDA receptor activity / dendritic spine organization / positive regulation of synapse assembly / RAF/MAP kinase cascade / NMDA selective glutamate receptor signaling pathway / beta-2 adrenergic receptor binding / Synaptic adhesion-like molecules / neuromuscular process controlling balance / neurotransmitter receptor localization to postsynaptic specialization membrane / neuron projection terminus / cortical cytoskeleton / AMPA glutamate receptor clustering / locomotory exploration behavior / AMPA glutamate receptor complex / Unblocking of NMDA receptors, glutamate binding and activation / regulation of neuronal synaptic plasticity / glutamate receptor binding / social behavior / kinesin binding / D1 dopamine receptor binding / positive regulation of synaptic transmission / regulation of postsynaptic membrane neurotransmitter receptor levels / excitatory synapse / ionotropic glutamate receptor binding / dendrite cytoplasm / positive regulation of excitatory postsynaptic potential / cell periphery / synaptic membrane / PDZ domain binding / neuromuscular junction / establishment of protein localization / cell-cell adhesion / cerebral cortex development / regulation of long-term neuronal synaptic plasticity / postsynaptic density membrane / kinase binding / cell-cell junction / synaptic vesicle / cell junction / nervous system development / positive regulation of cytosolic calcium ion concentration / protein-containing complex assembly / scaffold protein binding / protein phosphatase binding / dendritic spine / chemical synaptic transmission / postsynaptic membrane / neuron projection / postsynaptic density / postsynapse / signaling receptor binding / synapse / dendrite / protein kinase binding / protein-containing complex binding / glutamatergic synapse / endoplasmic reticulum / membrane / plasma membrane / cytoplasm / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method | SOLUTION NMR | ||||||
 Authors Authors | Tochio, H. / Hung, F. / Li, M. / Zhang, M. | ||||||
 Citation Citation |  Journal: J Mol Biol / Year: 2000 Journal: J Mol Biol / Year: 2000Title: Solution structure and backbone dynamics of the second PDZ domain of postsynaptic density-95. Authors: H Tochio / F Hung / M Li / D S Bredt / M Zhang /  Abstract: The second PDZ domain of postsynaptic density-95 (PSD-95 PDZ2) plays a critical role in coupling N-methyl-D-aspartate receptors to neuronal nitric oxide synthase (nNOS). In this work, the solution ...The second PDZ domain of postsynaptic density-95 (PSD-95 PDZ2) plays a critical role in coupling N-methyl-D-aspartate receptors to neuronal nitric oxide synthase (nNOS). In this work, the solution structure of PSD-95 PDZ2 was determined to high resolution by NMR spectroscopy. The structure of PSD-95 PDZ2 was compared in detail with that of alpha1-syntrophin PDZ domain, as the PDZ domains share similar target interaction properties. The interaction of the PSD-95 PDZ2 with a carboxyl-terminal peptide derived from a cytoplasmic protein CAPON was studied by NMR titration experiments. Complex formation between PSD-95 PDZ2 and the nNOS PDZ was modelled on the basis of the crystal structure of the alpha1-syntrophin PDZ/nNOS PDZ dimer. We found that the prolonged loop connecting the betaB and betaC strands of PSD-95 PDZ2 is likely to play a role in both the binding of the carboxyl-terminal peptide and the nNOS beta-finger. Finally, the backbone dynamics of the PSD-95 PDZ2 in the absence of bound peptide were studied using a model-free approach. The "GLGF"-loop and the loop connecting alphaB and betaF of the protein display some degree of flexibility in solution. The rest of the protein is rigid and lacks detectable slow time-scale (microseconds to milliseconds) motions. In particular, the loop connecting betaB and betaC loop adopts a well-defined, rigid structure in solution. It appears that the loop adopts a pre-aligned conformation for the PDZ domain to interact with its targets. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1qlc.cif.gz 1qlc.cif.gz | 552.7 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1qlc.ent.gz pdb1qlc.ent.gz | 462.1 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1qlc.json.gz 1qlc.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ql/1qlc https://data.pdbj.org/pub/pdb/validation_reports/ql/1qlc ftp://data.pdbj.org/pub/pdb/validation_reports/ql/1qlc ftp://data.pdbj.org/pub/pdb/validation_reports/ql/1qlc | HTTPS FTP |
|---|
-Related structure data
| Related structure data | |
|---|---|
| Similar structure data |
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| 1 |
| |||||||||
| NMR ensembles |
|
- Components
Components
| #1: Protein | Mass: 9982.496 Da / Num. of mol.: 1 / Fragment: THE SECOND PDZ DOMAIN Source method: isolated from a genetically manipulated source Source: (gene. exp.)   |
|---|
-Experimental details
-Experiment
| Experiment | Method: SOLUTION NMR |
|---|
- Sample preparation
Sample preparation
| Sample conditions | Ionic strength: 100 mM / pH: 6 / Pressure: 1 atm / Temperature: 303 K |
|---|---|
| Crystal grow | *PLUS Method: other / Details: NMR |
-NMR measurement
| NMR spectrometer | Type: Varian UNITY / Manufacturer: Varian / Model: UNITY / Field strength: 500 MHz |
|---|
- Processing
Processing
| NMR software | Name:  X-PLOR / Version: 3.8 / Developer: BRUNGER / Classification: refinement X-PLOR / Version: 3.8 / Developer: BRUNGER / Classification: refinement |
|---|---|
| Refinement | Software ordinal: 1 Details: TORSION ANGLE DYNAMICS SIMULATED ANNEALING REFINEMENT |
| NMR ensemble | Conformers submitted total number: 20 |
 Movie
Movie Controller
Controller













 PDBj
PDBj



