+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1oun | ||||||
|---|---|---|---|---|---|---|---|
| Title | CRYSTAL STRUCTURE OF NUCLEAR TRANSPORT FACTOR 2 (NTF2) | ||||||
 Components Components | NUCLEAR TRANSPORT FACTOR 2 | ||||||
 Keywords Keywords | TRANSPORT / NUCLEAR TRANSPORT PROTEIN | ||||||
| Function / homology |  Function and homology information Function and homology informationnegative regulation of vascular endothelial growth factor production / protein localization to nuclear pore / nuclear pore central transport channel / structural constituent of nuclear pore / nuclear inner membrane / nuclear import signal receptor activity / nuclear outer membrane / mRNA transport / protein export from nucleus / positive regulation of protein import into nucleus ...negative regulation of vascular endothelial growth factor production / protein localization to nuclear pore / nuclear pore central transport channel / structural constituent of nuclear pore / nuclear inner membrane / nuclear import signal receptor activity / nuclear outer membrane / mRNA transport / protein export from nucleus / positive regulation of protein import into nucleus / small GTPase binding / protein import into nucleus / nuclear membrane / nucleoplasm / identical protein binding / cytosol Similarity search - Function | ||||||
| Biological species |  | ||||||
| Method |  X-RAY DIFFRACTION / SINGLE ISOMORPHOUS REPLACEMENT / Resolution: 2.3 Å X-RAY DIFFRACTION / SINGLE ISOMORPHOUS REPLACEMENT / Resolution: 2.3 Å | ||||||
 Authors Authors | Bullock, T.L. / Stewart, M.J. | ||||||
 Citation Citation |  Journal: J.Mol.Biol. / Year: 1996 Journal: J.Mol.Biol. / Year: 1996Title: The 1.6 angstroms resolution crystal structure of nuclear transport factor 2 (NTF2). Authors: Bullock, T.L. / Clarkson, W.D. / Kent, H.M. / Stewart, M. #1:  Journal: Structure / Year: 1994 Journal: Structure / Year: 1994Title: Crystal Structure of Scytalone Dehydratase--A Disease Determinant of the Rice Pathogen, Magnaporthe Grisea Authors: Lundqvist, T. / Rice, J. / Hodge, C.N. / Basarab, G.S. / Pierce, J. / Lindqvist, Y. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1oun.cif.gz 1oun.cif.gz | 61.1 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1oun.ent.gz pdb1oun.ent.gz | 46 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1oun.json.gz 1oun.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/ou/1oun https://data.pdbj.org/pub/pdb/validation_reports/ou/1oun ftp://data.pdbj.org/pub/pdb/validation_reports/ou/1oun ftp://data.pdbj.org/pub/pdb/validation_reports/ou/1oun | HTTPS FTP |
|---|
-Related structure data
| Similar structure data |
|---|
- Links
Links
- Assembly
Assembly
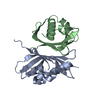
| Deposited unit | 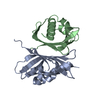
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 14491.425 Da / Num. of mol.: 2 Source method: isolated from a genetically manipulated source Source: (gene. exp.)   #2: Water | ChemComp-HOH / | |
|---|
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION / Number of used crystals: 1 X-RAY DIFFRACTION / Number of used crystals: 1 |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.48 Å3/Da / Density % sol: 47 % | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | pH: 4.5 / Details: pH 4.5 | ||||||||||||||||||||
| Crystal grow | *PLUS Method: vapor diffusion / PH range low: 5.5 / PH range high: 4.5 | ||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Diffraction | Mean temperature: 293 K |
|---|---|
| Diffraction source | Source:  ROTATING ANODE / Type: ELLIOTT GX-13 / Wavelength: 1.5418 ROTATING ANODE / Type: ELLIOTT GX-13 / Wavelength: 1.5418 |
| Detector | Type: MARRESEARCH / Detector: IMAGE PLATE / Date: Sep 1, 1995 |
| Radiation | Monochromatic (M) / Laue (L): M / Scattering type: x-ray |
| Radiation wavelength | Wavelength: 1.5418 Å / Relative weight: 1 |
| Reflection | Resolution: 2.3→100 Å / Num. obs: 12825 / % possible obs: 96.9 % / Observed criterion σ(I): 0 / Redundancy: 5.8 % / Rmerge(I) obs: 0.059 / Net I/σ(I): 9 |
| Reflection shell | Resolution: 2.3→2.42 Å / Redundancy: 5.7 % / Rmerge(I) obs: 0.175 / Mean I/σ(I) obs: 4.1 / % possible all: 94.7 |
| Reflection | *PLUS Redundancy: 5.4 % / Num. measured all: 90240 |
| Reflection shell | *PLUS % possible obs: 94.7 % |
- Processing
Processing
| Software |
| ||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Method to determine structure: SINGLE ISOMORPHOUS REPLACEMENT Resolution: 2.3→100 Å / σ(F): 0 Details: TNT LIBRARY RMS CO-ORDINATE SHIFT IN FINAL CYCLE (A) : 0.01
| ||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.3→100 Å
| ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||
| Software | *PLUS Name: TNT / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.158 | ||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS | ||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints | *PLUS
|
 Movie
Movie Controller
Controller












 PDBj
PDBj

