+ Open data
Open data
- Basic information
Basic information
| Entry | Database: PDB / ID: 1ohj | ||||||
|---|---|---|---|---|---|---|---|
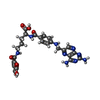
| Title | HUMAN DIHYDROFOLATE REDUCTASE, MONOCLINIC (P21) CRYSTAL FORM | ||||||
 Components Components | DIHYDROFOLATE REDUCTASE | ||||||
 Keywords Keywords | OXIDOREDUCTASE | ||||||
| Function / homology |  Function and homology information Function and homology informationregulation of removal of superoxide radicals / tetrahydrobiopterin biosynthetic process / Metabolism of folate and pterines / tetrahydrofolate metabolic process / response to methotrexate / sequence-specific mRNA binding / folic acid binding / axon regeneration / dihydrofolate metabolic process / G1/S-Specific Transcription ...regulation of removal of superoxide radicals / tetrahydrobiopterin biosynthetic process / Metabolism of folate and pterines / tetrahydrofolate metabolic process / response to methotrexate / sequence-specific mRNA binding / folic acid binding / axon regeneration / dihydrofolate metabolic process / G1/S-Specific Transcription / dihydrofolate reductase / dihydrofolate reductase activity / folic acid metabolic process / tetrahydrofolate biosynthetic process / NADPH binding / Tetrahydrobiopterin (BH4) synthesis, recycling, salvage and regulation / one-carbon metabolic process / mRNA regulatory element binding translation repressor activity / NADP binding / molecular adaptor activity / negative regulation of translation / mRNA binding / mitochondrion / cytosol Similarity search - Function | ||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | ||||||
| Method |  X-RAY DIFFRACTION / Resolution: 2.5 Å X-RAY DIFFRACTION / Resolution: 2.5 Å | ||||||
 Authors Authors | Cody, V. / Galitsky, N. / Luft, J.R. / Pangborn, W. | ||||||
 Citation Citation |  Journal: Biochemistry / Year: 1997 Journal: Biochemistry / Year: 1997Title: Comparison of two independent crystal structures of human dihydrofolate reductase ternary complexes reduced with nicotinamide adenine dinucleotide phosphate and the very tight-binding inhibitor PT523. Authors: Cody, V. / Galitsky, N. / Luft, J.R. / Pangborn, W. / Rosowsky, A. / Blakley, R.L. #1:  Journal: J.Biol.Chem. / Year: 1995 Journal: J.Biol.Chem. / Year: 1995Title: Methotrexate-Resistant Variants of Human Dihydrofolate Reductase with Substitutions of Leucine 22. Kinetics, Crystallography, and Potential as Selectable Markers Authors: Lewis, W.S. / Cody, V. / Galitsky, N. / Luft, J.R. / Pangborn, W. / Chunduru, S.K. / Spencer, H.T. / Appleman, J.R. / Blakley, R.L. #2:  Journal: J.Biol.Chem. / Year: 1994 Journal: J.Biol.Chem. / Year: 1994Title: Methotrexate-Resistant Variants of Human Dihydrofolate Reductase. Effects of Phe31 Substitutions Authors: Chunduru, S.K. / Cody, V. / Luft, J.R. / Pangborn, W. / Appleman, J.R. / Blakley, R.L. #3:  Journal: Anti-Cancer Drug Des. / Year: 1992 Journal: Anti-Cancer Drug Des. / Year: 1992Title: Crystal Structure Determination at 2.3 A of Recombinant Human Dihydrofolate Reductase Ternary Complex with Nadph and Methotrexate-Gamma-Tetrazole Authors: Cody, V. / Luft, J.R. / Ciszak, E. / Kalman, T.I. / Freisheim, J.H. | ||||||
| History |
|
- Structure visualization
Structure visualization
| Structure viewer | Molecule:  Molmil Molmil Jmol/JSmol Jmol/JSmol |
|---|
- Downloads & links
Downloads & links
- Download
Download
| PDBx/mmCIF format |  1ohj.cif.gz 1ohj.cif.gz | 53.8 KB | Display |  PDBx/mmCIF format PDBx/mmCIF format |
|---|---|---|---|---|
| PDB format |  pdb1ohj.ent.gz pdb1ohj.ent.gz | 38.3 KB | Display |  PDB format PDB format |
| PDBx/mmJSON format |  1ohj.json.gz 1ohj.json.gz | Tree view |  PDBx/mmJSON format PDBx/mmJSON format | |
| Others |  Other downloads Other downloads |
-Validation report
| Arichive directory |  https://data.pdbj.org/pub/pdb/validation_reports/oh/1ohj https://data.pdbj.org/pub/pdb/validation_reports/oh/1ohj ftp://data.pdbj.org/pub/pdb/validation_reports/oh/1ohj ftp://data.pdbj.org/pub/pdb/validation_reports/oh/1ohj | HTTPS FTP |
|---|
-Related structure data
- Links
Links
- Assembly
Assembly
| Deposited unit | 
| ||||||||
|---|---|---|---|---|---|---|---|---|---|
| 1 |
| ||||||||
| Unit cell |
|
- Components
Components
| #1: Protein | Mass: 21349.525 Da / Num. of mol.: 1 Source method: isolated from a genetically manipulated source Details: COMPLEXED WITH NADPH AND PT523 NA-(4-AMINO-4-DEOXYPTEROYL)-ND-HEMIPHTHALOYL-L-ORNITHINE Source: (gene. exp.)  Homo sapiens (human) / Gene: POTENTIAL / Production host: Homo sapiens (human) / Gene: POTENTIAL / Production host:  |
|---|---|
| #2: Chemical | ChemComp-NDP / |
| #3: Chemical | ChemComp-COP / |
| #4: Water | ChemComp-HOH / |
-Experimental details
-Experiment
| Experiment | Method:  X-RAY DIFFRACTION X-RAY DIFFRACTION |
|---|
- Sample preparation
Sample preparation
| Crystal | Density Matthews: 2.54 Å3/Da / Density % sol: 48 % | |||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Crystal grow | *PLUS Temperature: 20 ℃ / pH: 7 / Method: vapor diffusion, hanging drop | |||||||||||||||||||||||||||||||||||
| Components of the solutions | *PLUS
|
-Data collection
| Radiation | Scattering type: x-ray |
|---|---|
| Radiation wavelength | Relative weight: 1 |
| Reflection | Resolution: 2.5→8 Å / Num. obs: 5276 / % possible obs: 93.4 % / Observed criterion σ(I): 2 |
- Processing
Processing
| Software | Name: PROFFT / Classification: refinement | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Resolution: 2.5→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement step | Cycle: LAST / Resolution: 2.5→8 Å
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refine LS restraints |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Refinement | *PLUS Rfactor obs: 0.208 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Solvent computation | *PLUS | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Displacement parameters | *PLUS Biso mean: 16.86 Å2 |
 Movie
Movie Controller
Controller




















 PDBj
PDBj